Abstract
Each human body plays host to a microbial population which is both numerically vast (at around 1014 microbial cells) and phenomenally diverse (over 1,000 species). The majority of the microbial species in the gut have not been cultured but the application of culture-independent approaches for high throughput diversity and functionality analysis has allowed characterisation of the diverse microbial phylotypes present in health and disease.
Studies in monozygotic twins, showing that these retain highly similar microbiota decades after birth and initial colonisation, are strongly indicative that diversity of the microbiome is host-specific and affected by the genotype. Microbial diversity in the human body is reflected in both richness and evenness. Diversity increases steeply from birth reaching its highest point in early adulthood, before declining in older age. However, in healthy subjects there appears to be a core of microbial phylotypes which remains relatively stable over time.
Studies of individuals from diverse geopraphies suggest that clusters of intestinal bacterial groups tend to occur together, constituting ‘enterotypes’. So variation in intestinal microbiota is stratified rather than continuous and there may be a limited number of host/microbial states which respond differently to environmental influences. Exploration of enterotypes and functional groups may provide biomarkers for disease and insights into the potential for new treatments based on manipulation of the microbiome.
In health, the microbiota interact with host defences and exist in harmonious homeostasis which can then be disturbed by invading organisms or when ‘carpet bombing’ by antibiotics occurs. In a portion of individuals with infections, the disease will resolve itself without the need for antibiotics and microbial homeostasis with the host’s defences is restored. The administration of probiotics (live microorganisms which when administered in adequate amounts confer a health benefit on the host) represents an artificial way to enhance or stimulate these natural processes.
The study of innate mechanisms of antimicrobial defence on the skin, including the production of numerous antimicrobial peptides (AMPs), has shown an important role for skin commensal organisms. These organisms may produce AMPs, and also amplify the innate immune responses to pathogens by activating signalling pathways and processing host produced AMPs. Research continues into how to enhance and manipulate the role of commensal organisms on the skin. The challenges of skin infection (including diseases caused by multiply resistant organisms) and infestations remain considerable. The potential to re-colonise the skin to replace or reduce pathogens, and exploring the relationship between microbiota elsewhere and skin diseases are among a growing list of research targets.
Lactobacillus species are among the best known ‘beneficial’ bacterial members of the human microbiota. Of the approximately 120 species known, about 15 are known to occur in the human vagina. These organisms have multiple properties, including the production of lactic acid, hydrogen peroxide and bacteriocins, which render the vagina inhospitable to potential pathogens. Depletion of the of the normal Lactobacillus population and overgrowth of vaginal anaerobes, accompanied by the loss of normal vaginal acidity can lead to bacterial vaginosis – the commonest cause of abnormal vaginal discharge in women. Some vaginal anaerobes are associated with the formation of vaginal biofilms which serve to act as a reservoir of organisms which persists after standard antibiotic therapy of bacterial vaginosis and may help to account for the characteristically high relapse rate in the condition. Administration of Lactobacillus species both vaginally and orally have shown beneficial effects in the treatment of bacterial vaginosis and such treatments have an excellent overall safety record.
Candida albicans is a frequent coloniser of human skin and mucosal membranes, and is a normal part of the microbiota in the mouth, gut and vagina. Nevertheless Candida albicans is the most common fungal pathogen worldwide and is a leading cause of serious and often fatal nosocomial infections. What turns this organism from a commensal to a pathogen is a combination of increasing virulence in the organism and predisposing host factors that compromise immunity. There has been considerable research into the use of probioticLactobacillus spp. in vaginal candidiasis. Studies in reconstituted human epithelium and monolayer cell cultures have shown that L. rhamnosus GG can protect mucosa from damage caused by Candida albicans, and enhance the immune responses of mucosal surfaces. Such findings offer the promise that the use of such probiotic bacteria could provide new options for antifungal therapy.
Studies of changes of the human intestinal microbiota in health and disease are complicated by its size and diversity. The Alimentary Pharmabiotic Centre in Cork (Republic of Ireland) has the mission to ‘mine microbes for mankind’ and its work illustrates the potential benefits of understanding the gut microbiota. Work undertaken at the centre includes: mapping changes in the microbiota with age; studies of the interaction between the microbiota and the gut; potential interactions between the gut microbiota and the central nervous system; the potential for probiotics to act as anti-infectives including through the production of bacteriocins; and the characterisation of interactions between gut microbiota and bile acids which have important roles as signalling molecules and in immunity.
The important disease entity where the role of the gut microbiota appears to be central is the Irritable Bowel Syndrome (IBS). IBS patients show evidence of immune activation, impaired gut barrier function and abnormal gut microbiota. Studies with probiotics have shown that these organisms can exert anti-inflammatory effects in inflammatory bowel disease and may strengthen the gut barrier in IBS of the diarrhoea-predominant type. Formal randomised trials of probiotics in IBS show mixed results with limited benefit for some but not all.
Studies confirm that administered probiotics can survive and temporarily colonise the gut. They can also stimulate the numbers of other lactic acid bacilli in the gut, and reduce the numbers of pathogens. However consuming live organisms is not the only way to influence gut microbiota. Dietary prebiotics are selectively fermented ingredients that can change the composition and/or activity of the gastrointestinal microbiota in beneficial ways. Dietary components that reach the colon, and are available to influence the microbiota include poorly digestible carbohydrates, such as non-starch polysaccharides, resistant starch, non-digestible oligosaccharides (NDOs) and polyphenols. Mixtures of probiotic and prebiotic ingredients that can selectively stimulate growth or activity of health promoting bacteria have been termed ‘synbiotics’. All of these approaches can influence gut microbial ecology, mainly to increase bifidobacteria and lactobacilli, but metagenomic approaches may reveal wider effects. Characterising how these changes produce physiological benefits may enable broader use of these tactics in health and disease in the future.
The current status of probiotic products commercially available worldwide is less than ideal. Prevalent problems include misidentification of ingredient organisms and poor viability of probiotic microorganisms leading to inadequate shelf life. On occasions these problems mean that some commercially available products cannot be considered to meet the definition of a probiotic product. Given the potential benefits of manipulating the human microbiota for beneficial effects, there is a clear need for improved regulation of probiotics.
The potential importance of the human microbiota cannot be overstated. ‘We feed our microbes, they talk to us and we benefit. We just have to understand and then exploit this.’ (Willem de Vos).
IMPACT OF MICROBIOTA IN HEALTH AND DISEASE
Professor Willem M. de Vos, Wageningen University, The Netherlands.
The human body is home to a vast and complex realm of microbes. Each of us is host to around 1014 microbial cells (weighing about 2kg) representing more than 1,000 species1. Understanding the interaction of this microbiome with the host in both health and disease can provide important insights into how we can benefit from its presence.
In order to be tolerated by our bodies our intestinal flora are adapted for their environment and interact with our immune system2, binding to intestinal cells and mucus3 and eliciting specific immune responses2,4 (Figure 1).
Figure 1: Impact from Intestinal Interactions
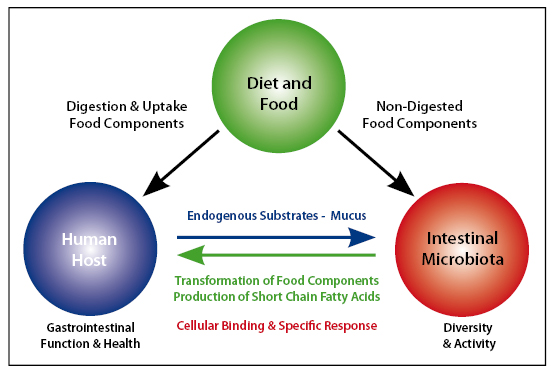
Numerically the microbiome dominates our bodies, dwarfing the amount of genetic material derived from our own cells (Figure 2).
Figure 2: Human GI Tract: Microbes Dominate Our Body
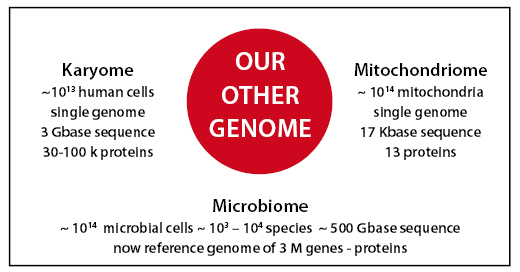
Adapted from Qin et al MetaHit Consortium.
A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464 (2010) 59-65
The great majority (up to about 80%) of these microbial species have not been cultured and this has led to applications of novel culture-independent approaches to provide a phylogenetic framework for the study of the more than 1000 different intestinal species. In recent years the use of techniques for high throughput diversity and functionality analysis of the gastrointestinal tract microbiota has allowed fundamental questions about about the role of microbes in the gut to be explored5,6,7 (Figure 3).
Figure 3: How to Learn More from the Gut
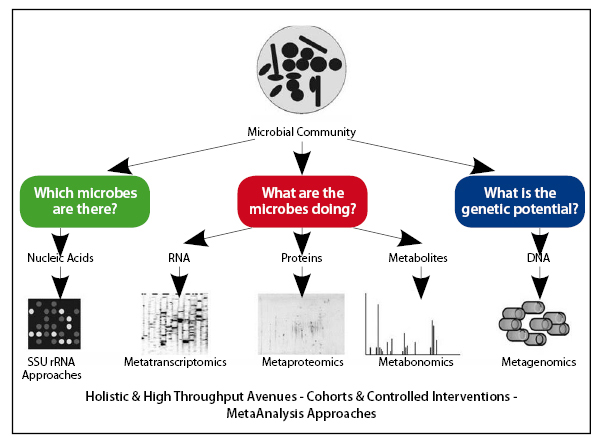
Reproduced from Gut, Zoetendal EG, M Rajilic-Stojanovic & WM de Vos. Vol 57, issue 11, pages 1605-15, 2008. With permission from BMJ Publishing Group Ltd.
The development of the human intestinal tract (HIT) chip, a phylogenetic microarray has allowed much more rapid analysis of the phylotypes represented in the abundant normal microbiota throughout life8. Comparative analysis with other methodologies such as pyrosequencing shows that this technique is robust and highly reproducible9.
Studies in monozygotic twins, showing that these retain highly similar microbiota decades after birth and intial colonisation is strongly indicative that diversity of the microbiome is host-specific and affected by the genotype10.
Microbial diversity in the human body shows both richness and eveness. Diversity increases steeply from birth reaching its highest point in early adulthood, before declining in older age11. Comparing the extremes of age shows marked differences in the composition of the microbiome with lower representation of butyrate producing species and an increased presence of potential pathogens in centenarians (Figure 4).
Figure 4: Marked Differences Between Young & Old (adapted from Biagi et al11)
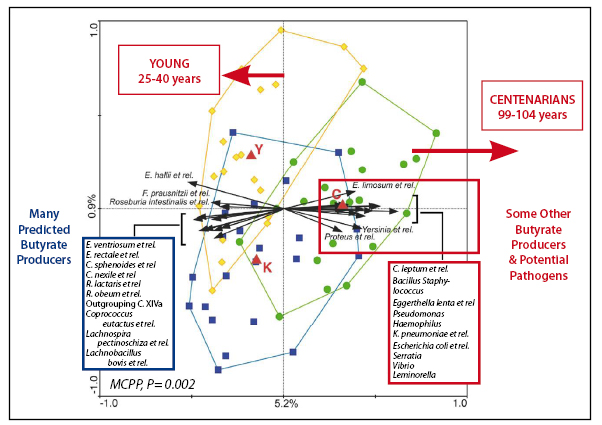
Diversity may be important and the composition of the microbiota may influence the health of babies from the earliest days of life. Babies crying because of intestinal colic showed significantly lower levels of microbial diversity than controls at 2 and 4 weeks. Representational Difference Analysis (RDA) analysis showed that 28% of this variance was explained by differences in a dozen microbial groups12.
It appears that there is a common core of approximately 400 microbial phylotypes in normal healthy subjects. Sequential sampling of individuals suggests that each individual has an individual core of phylotypes which remains stable over time13. Individual variation may explain the bulk of variation in populations with even extreme dietary manipulation (e.g. a very low caloric diet) producing relatively low levels of variation (60% vs 10% respectively).
Studies of sequenced faecal metagenomes of individuals from different countries has identified specific clusters of co-occurring bacterial groups, termed enterotypes. Confirmation of these enterotypes in larger cohorts, indicates that variation in intestinal microbiota is generally stratified rather than continuous14. This, in turn, suggests the existence of a limited number of host–microbial symbiotic states that might respond differently to diet and drug intake. Clustering of phylotypes in enterotypes and functional groups has enabled the search for early biomarkers of disease as well as providing new insights into the potential for novel therapies based on manipulation of the microbiome.
EXPERIMENTAL MICROBIAL THERAPIES
When a clinical condition is clearly caused by disturbance of the normal gut microbiota, such as in C.difficile Infections (CDI), one possibility is to restore the normal intestinal microbiota through ‘transplantation’ of normal donor colonic or duodenal infusions. This ‘faecal transplantation’ has been tried in hundreds of cases over the last 50 years and seems to be characterized by a high degree of success15.
Recent studies have allowed the application of modern analysis to characterise the changes of microbiota on transplantation16. Deep and high throughput analysis of the microbiota for several months after transplantation using the HIT Chip confirmed the correction of the microbiota17:
• Initial low diversity in patients with CDI
• correction of low diversity by the transplantations
• stability of the newly established diverse community
• a clear shift in the microbiota of the recipients towards the donor signature, correcting the effect of the patients’ gram-negative pathogens.
IMPACT OF GI TRACT MICROBES ON OBESITY
Faecal transplantation studies in mice show that transferring the microbiota from lean and fat mice to germ-free mice induces greater weight gain in those receiving the ‘fat’ microbiota. The discovery of this obesity-associated microbiome18, raises the possibility of using transplanted microbiota to influence metabolic processes in humans.
In a controlled study (autologous transplantation) examining the effect of faecal transplantation from lean subjects to those with metabolic syndrome, results showed significant improvement of whole body insulin sensitivity at 6 weeks19.
SUMMARY AND CONCLUSION
Recent years have witnessed a microbiota revolution as we have utilised methodologies that allow us to study the diversity, enterotypes, and functional signatures of the intestinal microbiota. This has enabled the recognition of early biomarkers, provided new insights into diseases, and helped to develop novel therapies. Importantly it has also allowed us to focus on human rather than model animals.
The potential importance of the human microbiota cannot be overstated. Put simply, we feed our microbes, they talk to us and we benefit. We just have to understand and then exploit this.
References
- Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J,Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto JM, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Doré J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J; MetaHIT Consortium, Bork P, Ehrlich SD, Wang J. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010 Mar 4;464(7285):59-65.
- van Baarlen P, Troost FJ, van Hemert S, van der Meer C, de Vos WM, de Groot PJ, Hooiveld GJ, Brummer RJ, Kleerebezem M. Differential NF-kappaB pathways induction by Lactobacillus plantarum in the duodenum of healthy humans correlating with immune tolerance. Proc Natl Acad Sci USA. 2009 Feb 17;106(7):2371-6. Epub 2009 Feb 3.
- Konstantinov SR, Smidt H, de Vos WM, Bruijns SC, Singh SK, Valence F, Molle D, Lortal S, Altermann E, Klaenhammer TR, van Kooyk Y. S layer protein A of Lactobacillus acidophilus NCFM regulates immature dendritic cell and T cell functions. Proc Natl Acad Sci USA. 2008 Dec 9;105(49):19474-9. Epub 2008 Dec 1.
- Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx AP, Lebeer S, De Keersmaecker SC, Vanderleyden J, Hämäläinen T, Laukkanen S, Salovuori N, Ritari J, Alatalo E, Korpela R, Mattila-Sandholm T, Lassig A, Hatakka K, Kinnunen KT, Karjalainen H, Saxelin M, Laakso K, Surakka A, Palva A, Salusjärvi T, Auvinen P, de Vos WM. Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci U S A. 2009 Oct 6;106(40):17193-8. Epub 2009 Sep 17
- Lay C, Rigottier-Gois L, Holmstrøm K, Rajilic M, Vaughan EE, de Vos WM, Collins MD, Thiel R, Namsolleck P, Blaut M, Doré J. Colonic microbiota signatures across five northern European countries. Appl Environ Microbiol. 2005 Jul;71(7):4153-5.
- Zoetendal EG, Vaughan EE, de Vos WM. A microbial world within us. Mol Microbiol. 2006 Mar;59(6):1639-50. Review.
- Zoetendal EG, Rajilic-Stojanovic M, de Vos WM. High-throughput diversity and functionality analysis of the gastrointestinal tract microbiota. Gut. 2008 Nov;57(11):1605-15. Review.
- Rajilic-Stojanovic M, Heilig HG, Molenaar D, Kajander K, Surakka A, Smidt H, de Vos WM. Development and application of the human intestinal tract chip, a phylogenetic microarray: analysis of universally conserved phylotypes in the abundant microbiota of young and elderly adults. Environ Microbiol. 2009 Jul;11(7):1736-51. Epub 2009 Mar 11.
- Claesson MJ, O’Sullivan O, Wang Q, Nikkilä J, Marchesi JR, Smidt H, de Vos WM, Ross RP, O’Toole PW. Comparative analysis of pyrosequencing and a phylogenetic microarray for exploring microbial community structures in the human distal intestine. PLoS One. 2009 Aug 20;4(8):e6669.
- Zoetandal EG, Akkermans ADL, Akkermans-van Vliet WM, de Visser JA, de Vos WM. The host genotype affects the bacterial community in the human gastrointestinal tract. Microb. Ecol. Health Dis. 2001 13, 129-134.
- Biagi E, Nylund L, Candela M, Ostan R, Bucci L, Pini E, Nikkïla J, Monti D, Satokari R, Franceschi C, Brigidi P, de Vos W. Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS One. 2010 May 17;5(5):e10667. Erratum in: PLoS One. 2010;5(6). doi: 10.1371/annotation/df45912f-d15c-44ab-8312-e7ec0607604d.
- de Weerth C, Fuentes S, Puylaert P, de Vos WM. Intestinal microbiota of infants with colic: development and specific signatures. Paediatrics 2012 (in press).
- Jalanka-Tuovinen J, Salonen A, Nikkilä J, Immonen O, Kekkonen R, Lahti L, Palva A, de Vos WM. Intestinal microbiota in healthy adults: temporal analysis reveals individual and common core and relation to intestinal symptoms. PLoS One. 2011;6(7):e23035. Epub 2011 Jul 28.
- Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR, Fernandes GR, Tap J, Bruls T, Batto JM, Bertalan M, Borruel N, Casellas F, Fernandez L, Gautier L, Hansen T, Hattori M, Hayashi T, Kleerebezem M, Kurokawa K, Leclerc M, Levenez F, Manichanh C, Nielsen HB, Nielsen T, Pons N, Poulain J, Qin J, Sicheritz-Ponten T, Tims S, Torrents D, Ugarte E, Zoetendal EG, Wang J, Guarner F, Pedersen O, de Vos WM, Brunak S, Doré J; MetaHIT Consortium,Weissenbach J, Ehrlich SD, Bork P. Enterotypes of the human gut microbiome. Nature. 2011 May 12;473(7346):174-80. Epub 2011 Apr 20. Erratum in: Nature. 2011 Jun 30;474(7353):666.
- van Nood E, Speelman P, Kuijper EJ, Keller JJ. Struggling with recurrent Clostridium difficile infections: is donor faeces the solution? Euro Surveill. 2009 Aug 27;14(34). pii: 19316. Review.
- Khoruts A, Dicksved J, Jansson JK, Sadowsky MJ. Changes in the composition of the human fecal microbiome after bacteriotherapy for recurrent Clostridium difficile-associated diarrhea. J Clin Gastroenterol. 2010 May-Jun;44(5):354-60.
- van Nood E, Vrieze A, Nieuwdorp M, Fuentes S, Zoetendal EG, de Vos WM, Visser CE, Kuijper EJ, Bartelsman JF, Tijssen JGP, Speelman P, Dijkgraaf MGW, Keller JJ. Donor Feces Infusion for Recurrent Clostridium difficile Infections. Submitted for publication.
- Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006 Dec 21;444(7122):1027-31.
- Vrieze A, van Nood E, Holleman F, Salojärvi J, Kootte RS, Bartelsman JF, Dallinga-Thie GM, Ackermans MT, Serlie MJ, Oozeer R, Derrien M, Druesne A, van Hylckama Vlieg JE, Bloks VW, Groen AK, Heilig HG, Zoetendal EG, Stroes ES, de Vos WM, Hoekstra JB, Nieuwdorp M. Transfer of Intestinal Microbiota from Lean Donors Increases Insulin Sensitivity in Subjects with Metabolic Syndrome. Gastroenterology. 2012 October 143;4:913–916.e7.
ANALYTICS OF THE HUMAN MICROBIOTA
Professor Lars Engstrand, Karolinska Institute, Stockholm.
The gastrointestinal tract shows progressive colonization from the mouth to the colon with markedly different proportions of bacterial phyla present depending on the site of sampling. Humans are born sterile, but during birth a microbial population starts to establish in the gut. At first, the composition is very variable, with mainly facultative anaerobic or aerobic bacteria but after approximately 2 years it stabilizes. Composition of the large bowel is then mainly obligate anaerobes, Bacteroides and Firmicutes, such as clostridia.
The gut microbiota is affected by a number of factors including: mode of delivery at birth (vaginal or caesarean section), host genotype and lifestyle factors (including dietary habits). Systemic antibiotics can have long term effects on microbial composition of the gut and can disrupt some bacterial groups for several years.
The characterization of the human microbiota employs many tools and has given rise to entire fields of study (Figure 1).
Figure 1
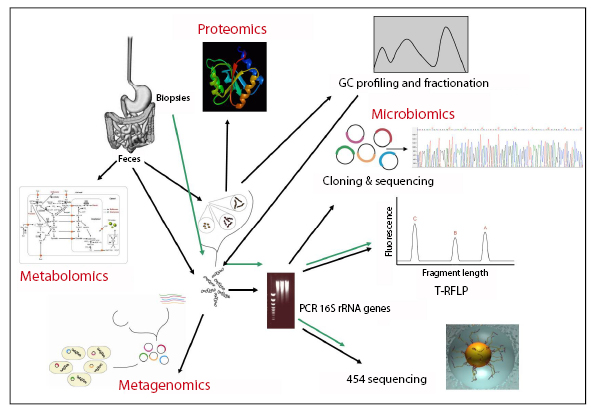
The next generation of gene sequencing technology will enable enormous increases in the speed of sequencing at greatly reduced cost. Combination of 454-pyrosequencing of a hyper-variable region of the 16S rRNA gene in combination with sample specific barcode sequences, enables parallel in-depth analysis of hundreds of samples with limited sample processing1.
Modeling demonstrates that the method correctly describes microbial communities down to phylotypes below the genus level. Determining the microbial composition in patients and healthy controls using this technology may also provide novel therapeutic targets.
Figure 2: heatmap of bacterial diversity
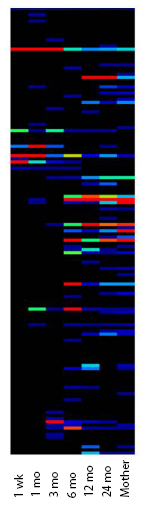

Figure 2 shows a ‘heatmap’ of bacterial diversity illustrating the increase in diversity in samples from infants at different ages compared to their mother.
Figure 3: Outcomes of H. pylori infection
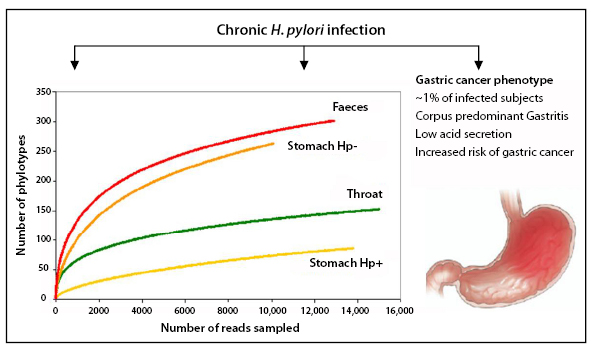
Figure 3 shows the number of phylotypes sampled from different sites in the gastrointestinal tract as a function of the number of reads. Illustrated here is the impact of Helicobacter pyloriinfection.
Figure 4: Inflammatory Bowel Disease in twins (T-RFLP)

Using these techniques we can compare the microbiota of subjects with and without diseases and study e.g. the effects of host genotypes on disease. Figure 4 shows the T-RFLP profiles for a healthy twin pair (left), a discordant twin pair (middle) and a concordant twin pair (right). The size of the pie slice represent relative abundance value. Black, blue, green, red and purple TRFs are also represented with the same color coding for Bacteroides sp. primers.
We have also been able to show the changes in diversity over time in the microbiota of infants whether they were born vaginally or by caesarean section. The genus abundance over time shows clear differences at early time points which largely disappear at 24 months under the influence of external factors such as feeding.
These techniques have also been used to study the gut bacteria of Ötzi – the iceman – an individual who lived in 3300 BC and was discovered well preserved in ice 5 millenia later. Sampling from various sites in the gastrointestinal tract yielded an impressive list of pathogens and associated virulence factors.
SUMMARY
The use of 454-pyrosequencing of a hyper-variable region of the 16S rRNA gene in combination with sample specific barcode sequences has great potential to help in our understanding of the human microbiota in health and disease. Pilot studies are already underway in a number of diseases and areas of interest including:
• Bacterial vaginosis
• Psoriasis
• Paradontitis, neutropenia
• Ulcer and gastritis
• Neonatal necrotizing entercolitis
• Probiotics, Allergy, Birth.
ADDITIONAL READING
Pflughoeft KJ, Versalovic J. Human microbiome in health and disease. Annu Rev Pathol. 2012;7:99-122.
Maccaferri S, Biagi E, Brigidi P. Metagenomics: key to human gut microbiota.Dig Dis. 2011;29(6):525-30.
Jeffery IB, Quigley EM, Ohman L, Simrén M, O’Toole PW. The microbiota link to Irritable Bowel Syndrome: An emerging story. Gut Microbes. 2012 Nov 1;3(6). [Epub ahead of print]
Rogers GB, Bruce KD. Next-generation sequencing in the analysis of human microbiota: essential considerations for clinical application. Mol Diagn Ther. 2010 Dec 1;14(6):343-50.
Chen W, Liu F, Ling Z, Tong X, Xiang C. Human intestinal lumen and mucosa-associated microbiota in patients with colorectal cancer. PLoS One. 2012;7(6):e39743.
References
- Andersson AF, Lindberg M, Jakobsson H, Bäckhed F, Nyrén P, Engstrand L. Comparative analysis of human gut microbiota by barcoded pyrosequencing. PLoS One. 2008 Jul 30;3(7):e2836. .
STABILITY AND QUALITY OF COMMERCIALLY AVAILABLE PRODUCTS
Professor Lorenzo Drago, Lab. of Clinical Chemistry and Microbiology, IRCCS Galeazzi Hospital – Dept of Biomedical Sciences for Health, University of Milan, Italy.
Probiotics are live microorganisms which, when administered in adequate amounts, confer a health benefit on the host1.
Probiotics set a number of challenges for micriobiologists, namely:
Terminology
Defining the terminology associated with probiotics
Guidelines
Defining the characteristics of probiotics according to guidelines
Methodology
Developing new tools to evaluate preclinical and clinical efficacy or to better understand the mechanisms of action.
Safety
Monitoring of issues (i.e. genetic stability, antibiotic-resistance, local adverse effects) occurring over time.
In the last 2 decades (from June 1989 to March 2011) 6806 studies of probiotics have been published in indexed journals most dealing with efficacy but few addressing the quality of marketed probiotic products. Over approximately the same period there has been an evolution in the therapeutic areas where probiotics have been claimed to offer benefit. In the early 1990s these products were mainly promoted for diarrhoea, but in the last decade this has been generalised to intestinal dysfunction and irritable bowel syndrome with atopic diseases joining the list in 2008.
Probiotics vary greatly in popularity between countries: in 2009 in Italy there were 206 manufacturers of 1609 products of this type. In some cases, the sales of individual products seem inversely proportional to the amount of scientific literature generated about them.
The Guidelines drafted in 2002 by FAO/WHO1 proposed important guidance for identifying and characterizing organisms to species level, assessing safety and conducting clinical studies of efficacy.
The first of these is important since different strains of the same species may act in different ways and produce different results. For this reason it is absolutely essential to identify the strain of organism to be used as a probiotic and there are well established ways to do this by genotyping. However it has been known for some time that probiotic supplements may not contain what they are promoted to contain, may be of deficient microbiological quality, and may make misleading health claims2,3.
Some of the more frequent misidentifications of ingredient organisms include:
L. crispatus instead of L. acidophilus
L. gasseri instead of L. acidophilus
L. paracasei instead of L. casei or L. jughurti
and even Streptococcus sanguis instead of L. Acidophilus
Some probiotic organisms have been given made up names, presumably for marketing purposes, including: Bifidus actiregularis, Lactobacillus immunitass, Bifidus attivo essensis.
One important problem for manufacturers to overcome is the need for sufficient viable organisms to be delivered in the product, since a large load may be needed to survive gastric acid exposure. This consideration leads to recommendation to take doses of the order of 108 – 1011 colony forming units (cfu) per dose, one to several times per day.
Studies of probiotic products commercialised in Italy have consistently revealed deficiencies in their probiotic attributes4,5. Lactobacillus sporogenes, or Bacillus coagulans, as it should be correctly classified, represents the archetypal misidentified probiotic. Bacillus probiotics could offer some advantages, such as low cost of production processes, ease of preparation, resistance to production process and extended shelf life over a wide range of temperatures. Although the use of L. sporogenes spores as a probiotic has increased in recent years, clinical evidence of its benefits are limited to only a few studies involving small patient populations6.
More recently we conducted a study to determine if products available in the USA market in 2009 were correctly labeled in terms of quantity of viable bacteria, identification of species and cross contamination by species not on the label. Disturbingly, we found that only 4 of 13 products (31%) were in accordance with label claims. Furthermore 6/13 (46%) contained less viable bacteria than claimed, 5/13 (38%) had at least one species missing, and 7/13 (54%) contained contaminant organisms (Bacillus spp., Staphylococcus spp., Enterococcus faecium, molds)7.
In a similar, as yet unpublished study with 24 commercial products available in the Italian market in 2010, we found: 15/24 (62.5%) products in accordance with the label; 9/24 (37.5%) containing less viable bacteria than declared (regarding one species at least); 4/24 (17%) with at least one species missing; and 2/24 (8%) containing contaminantEnterococcus faecium.
Nonetheless, and considering their widespread use, probiotics have not been associated with many significant safety issues. There have been no deaths apparently due to probiotics (except for one study with acute pancreatitis patients)8. With regard to Lactobacillusbacteremias there have been 180 cases in 30 years and 69 cases of endocarditis.
There have been case reports of outbreaks caused by S. cerevesiae and S. Boulardii9,10,11.
One safety question that has arisen is ‘Can probiotics transfer antimicrobial resistance to other bacteria?’. From this several subsidiary questions arise:
• Is co-administration of antibiotics and probiotics appropriate ?
• Can a probiotic develop resistance?
• Can resistance become stable?
• Can a probiotic transfer resistance?
We have shown resistance to macrolides and/or tetracycline in 14 and 4 isolates respectively of 40 isolates of Lactobacillus acidophilus, Lactobacillus plantarum, Lactobacillus crispatus, and Lactobacillus casei isolated from faeces of apparently healthy volunteers. Serial exposure to antibiotics led to selection of resistant mutants. However, acquired resistance was rather unstable and was lost after subcultures in antibiotic-free medium in most mutants12. These findings lead us to recommend that since lactobacilli are often used as probiotics, their ability to acquire resistance should be evaluated for isolates that may be candidates to be included in probiotic based products.
In an unpublished study we examined the presence of antibiotic resistance in Italian probiotics. In 13 commercial products, containing 21 isolated strains we found:
• 12/13 (92%) products containing at least one strain showing resistance to at least one antibiotic
• 10/13 (77%) products containing at least one strain harbouring known resistance genes
And among the isolates:
• 5/21 (24%) erythromycin-resistant strains
• 2/21 (9,5%) tetracycline-resistant strains
• 14/21 (67%) gentamicin-resistant strains
• 0/21 (0%) penicillin-resistant strains
SUMMARY
Strain identification still remains a worldwide problem. Some products are misnamed and not correctly identified. A combination of proper molecular methods seems the most suitable approach to this problem. Viability of probiotic microorganisms is essential, but some marketed products do not possess a suitable shelf-life. It is possible that resistant genes may be spread via some probiotics. It is apparent that some commercially produces products should not be included on the list of true probiotics. There are many “fancy names” and much “mislabelling” and there is a clear need to improve regulatory issues.
References
- Report of a Joint FAO/WHO Expert Consultation on Evaluation of Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria (October 2001). “Health and Nutritional Properties of Probiotics in Food including Powder Milk with Live Lactic Acid Bacteria”. Food and Agriculture Organization of the United Nations, World Health Organization. Accessed May 4th 2012
Reference Link - Hamilton-Miller JM, Shah S, Smith CT. “Probiotic” remedies are not what they seem. BMJ. 1996 Jan 6;312(7022):55-6.
- Hamilton-Miller JM, Shah S. Deficiencies in microbiological quality and labelling of probiotic supplements. Int J Food Microbiol. 2002 Jan 30;72(1-2):175-6.
- Drago L, De Vecchi E, Nicola L, Colombo A, Gismondo MR. Microbiological evaluation of commercial probiotic products available in Italy. J Chemother. 2004 Oct;16(5):463-7.
- De Vecchi E, Nicola L, Zanini S, Drago L. In vitro screening of probiotic characteristics of some italian products. J Chemother. 2008 Jun;20(3):341-7.
- Drago L, De Vecchi E. Should Lactobacillus sporogenes and Bacillus coagulans have a future? J Chemother. 2009 Aug;21(4):371-7. Review.
- Drago L, Rodighiero V, Celeste T, Rovetto L, De Vecchi E. Microbiological evaluation of commercial probiotic products available in the USA in 2009. J Chemother. 2010 Dec;22(6):373-7.
- Besselink MG et al. Probiotic prophylaxis in predicted severe acute pancreatitis: a randomised, double-blind, placebo-controlled trial. Lancet 2008 371: 651–659.
- Thygesen JB, Glerup H, Tarp B. Saccharomyces boulardii fungemia caused by treatment with a probioticum. BMJ Case Rep. 2012 Mar 27;2012.
- Enache-Angoulvant A, Hennequin C. Invasive Saccharomyces infection: a comprehensive review. Clin Infect Dis. 2005 Dec 1;41(11):1559-68.
- Cherifi S, Robberecht J, Miendje Y. Saccharomyces cerevisiae fungemia in an elderly patient with Clostridium difficile colitis. Acta Clin Belg. 2004 Jul-Aug;59(4):223-4.)
- Drago L, Mattina R, Nicola L, Rodighiero V, De Vecchi E. Macrolide resistance and in vitro selection of resistance to antibiotics in Lactobacillus isolates. J Microbiol. 2011 Aug;49(4):651-6. Epub 2011 Sep 2.
MODES OF ACTION OF MICROBIOTA RESTORATION
Dr Gregor Reid, Lawson Health Research Institute, University of Western Ontario, London, Canada.
Inside every human, potentially pathogenic and non-pathogenic organisms can be found. When infection occurs, it can be due to an external source or by propagation of the person’s own organisms. Antibiotics and other chemicals can disturb the microbiota by essentially carpet bombing a niche and killing pathogens as well as so-called beneficial organisms. It takes many weeks for homeostasis to recover. In a portion of patients with infection, such as in the bladder, vagina or respiratory tract, infection can self-resolve without the intervention of antibiotics. This, and the damage caused by antibiotics, has lead to a search for ways to understand and restore microbial homeostasis more quickly.
Probiotics are “live microorganisms which when administered in adequate amounts confer a health benefit on the host”. Strains of probiotics have been examined for a role in preventing and managing infections, especially in the gut and vagina.
A key factor in applying probiotic interventions is to understand how the microbiome develops from the moment of birth, and thereby identifying what strains might foster a niche dominated by bacteria that benefit the host.
The assembly of the human infant gut microbiome is characterised by an increase in phylogenetic diversity over time. However the abundance of major taxonomic groups shows abrupt shifts corresponding to life events in the infant such as dietary changes from milk to solid food, with the functional genes present reflecting the foods to which the gut is exposed1.
So how might probiotics be used to affect the microbiota of the gut and vagina? McNulty et al.2 found ingestion of probiotic yogurt did not result in the organisms taking up residence in the human or mouse gut, nor marked alteration in the composition of the host gut microbiota, but it did alter metabolic pathways in the resident microbiota, particularly those related to carbohydrate processing. In other words, it had an effect on the host.
The study involved faecal sampling from adult female monozygotic twin pairs after consumption of a commercially available fermented milk product (FMP) containing a consortium of Bifidobacterium animalis subsp. lactis, two strains of Lactobacillus delbrueckiisubsp. bulgaricus, Lactococcus lactis subsp. cremoris, and Streptococcus thermophilus. There were no significant changes in bacterial species composition or in the proportional representation of genes encoding known enzymes observed in the faeces of the humans consuming the FMP. However the metatranscriptome exhibited significant changes, including most prominently those related to plant polysaccharide metabolism.
In other studies, Jacques Ravel’s group characterized the vaginal microbiome of 396 asymptomatic North American reproductive-age women who represented four ethnic groups (white, black, Hispanic, and Asian)3. Species composition was determined by pyrosequencing of barcoded 16S rRNA genes. The resident bacterial communities clustered into five groups: four were dominated by Lactobacillus iners, L. crispatus, L. gasseri, or L. jensenii, whereas the fifth had lower proportions of lactic acid bacteria and higher proportions of strictly anaerobic organisms, indicating that a key ecological function, the production of lactic acid, seems to be conserved in all communities. The proportions of each community group varied among the four ethnic groups, and these differences were statistically significant. This study indicates that the inherent differences within and between women in different ethnic groups may govern their risk of diseases including bacterial vaginosis (BV). But, it remains to be determined why ethnicity per se would cause this, or whether particular dietary intakes or sexual practices have over-riding influence.
Longitudinal studies of women show changes in the vaginal microbiota over time in individuals, largely coincident with menstruation, but there are differences between individuals, albeit generally lactobacilli are dominant in healthy women. In a study by Hummelen et al.4, four different forms of BV were identified based upon abundance of different pathogens. These differences may affect the success of the treatment options that were developed mostly to eradicate Gardnerella, not the other infecting strains. Lactobacilli can be present in low numbers during BV, and depending on the species, this can influence recovery of the microbiota.
A study of Lactobacillus crispatus CTV-05 administered vaginally after topical antibiotic treatment of BV showed that vaginal concentration of certain BV-associated bacteria (Gardnerella vaginalis and Atopobium spp.) and the absence of endogenous L. crispatus at enrollment predicted colonization with the probiotic lactobacilli5.
Placebo controlled clinical trials have confirmed that probiotic Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14 improve outcomes when used during and after single dose antimicrobial treatments for both BV and vulvovaginal candidiasis6-9. Why these combinations work is the subject of considerable research. Recent work by Kohler et al.10indicates that lactobacilli repress Candida genes involved in biofilm formation and fluconazole resistance and this may help explain the success of conjoint treatment with the probiotic and the antibiotic in eradicating the fungi.
Lactobacilli have a number of properties, including production of surfactants, hydrogen peroxide and bacteriocins, that are relevant to resisting pathogens (figure 1).
Figure 1

Reproduced by permission from Macmillan Publishers Ltd: Nature Reviews Microbiology.
Reid et al. 2011 Jan; 9(1):27-38. Epub 2010 Nov 29
The vagina may be subject to bacterial overgrowth of two distinct types. Aerobic vaginitis (AV) is characterized by inflammation, yellow discharge, and vaginal dyspareunia11. In this condition, group B streptococci, Escherichia coli, Staphylococcus aureus and Trichomonas vaginalis are frequently cultured. While BV is associated with an overgrowth of vaginal anaerobes and in some cases an offensive, fishy-smelling discharge, AV is an inflammatory condition that leads to high production of cytokines in the vaginal fluid.
In combatting bacterial overgrowth caused by aerobic or anaerobic bacteria, vaginal lactobacilli may be important in disrupting bacterial biofilms. The best known biofilm former,Pseudomonas aeruginosa, employs an intercellular communication system, known as quorum sensing (QS) to coordinate the expression of tissue-damaging factors. The QS system controls the production of different virulence factors. The results of real time quantitative polymerase chain reaction (PCR) have shown that in all P. aeruginosa strains grown in the presence of probiotic culture sterile filtrates, the level of QS genes expression was reduced comparatively with those from control cultures12. In the case of Lactobacillus iners, quorum sensing may help it adapt to an infected state, and form a nidus to recovery in some women.
Bacteriocins are molecules that inhibit the growth of like-species, but they can also act as quorum sensing agents. Scientists from the Alimentary Pharmabiotic Centre in Cork have demonstrated that Lactobacillus salivarius UCC118, a probiotic strain of human origin, produces a bacteriocin in vivo that can significantly protect mice against infection with the invasive foodborne pathogen Listeria monocytogenes13. This illustrates another mechanism whereby infection can be averted or overcome.
Indigenous or probiotic lactobacilli (Figure 2) may also help maintain the barrier functions of epithelial tight junctions. These linkages can be compromised by inflammation and pathogens14, inducing recruitment of macrophages and other host defenses to the site, leading to vaginal discharge.
Figure 2

Reproduced by permission from Macmillan Publishers Ltd: Nature Reviews Microbiology.
Reid et al. 2011 Jan; 9(1):27-38. Epub 2010 Nov 29
Figure 3

Reproduced by permission from Macmillan Publishers Ltd: Nature Reviews Microbiology.
Reid et al. 2011 Jan; 9(1):27-38. Epub 2010 Nov 29
LACTOBACILLI SIGNALLING SYSTEMS (FIGURE 3)
Some strains of Lactobacillus have signalling systems involved in downregulating toxin production in pathogenetic bacteria. A recent study shows that human vaginal isolateLactobacillus reuteri RC-14 produces small signaling molecules that are able to interfere with the staphylococcal quorum-sensing system agr, a key regulator of virulence genes, and repress the expression of TSST-1 in S. aureus MN815. This provides an example of one mechanism by which interspecies cell-to-cell communication between endogenous or probiotic strains and pathogens might attenuate virulence factor production by the latter.
Lactobacillus reuteri RC-14 can also disable a potent S. aureus exotoxin, toxic shock syndrome toxin (TSST-1)16. Intriguingly, vaginal swabs from women with BV and from normal women appear to suppress toxin production from S. aureus MN8 (manuscript in preparation).
Using atomic force microscopy, Younes et al16 have shown that the adhesion force of L. crispatus 33820 with S. aureus MN8 is greater than for the pathogen and the surface, indicating that the staphylococci are more likely to adhere to the lactobacilli than to remain on the surface. Notably, attachment occurs on contact and bond maturation occurs within a mere 120 seconds of physical contact between the lactobacilli and staphylococci. Co-aggregation is now one of the recognized mechanisms through which lactobacilli can exert their probiotic effects to create a hostile micro-environment around a pathogen.
Modulation of host immunity by non-pathogens might maintain immune homeostasis and preserve the microbiota composition and structure. This provides another potential mechanism by which probiotic strains might influence host defences. There is some evidence that lactobacilli also enhance local immunity. Human, animal and in vitro studies have shown that L. rhamnosus GR-1 down-regulates TNFα, PGE and Cox-2 inflammatory mediators and up-regulates IL-10, G-CSF, Treg cells, and antimicrobial IL-8, vaginal defensins α1, ß2, SLPI, NOD2, TLR2, TLR9, CD4 and elafin in HIV patients17-20.
Figure 4: The Interactome: The Toolbox is Filling Up (adapted from Bisanz and Reid21)

New techniques (e.g. meta-transcriptional and metabolomic profiling) will increasingly enable us to answer questions about what happens in the ‘Interactome’ between the human host, the microbiota and probiotic strains21 (figure 4).
SUMMARY
There are many cases in which infections resolve without intervention, likely due to host defenses and the microbiome’s ability to restore homeostasis. Probiotics are an artificial way to simulate or enhance these natural processes. We now have the tools and probiotic strains to demonstrate that microbiota restoration is possible on the skin, gut and vagina.
References
- Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R, Angenent LT, Ley RE. Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci U S A. 2011 Mar 15;108 Suppl 1:4578-85.
- McNulty NP, Yatsunenko T, Hsiao A, Faith JJ, Muegge BD, Goodman AL, Henrissat B, Oozeer R, Cools-Portier S, Gobert G, Chervaux C, Knights D, Lozupone CA, Knight R, Duncan AE, Bain JR, Muehlbauer MJ, Newgard CB, Heath AC, Gordon JI. The impact of a consortium of fermented milk strains on the gut microbiome of gnotobiotic mice and monozygotic twins. Sci Transl Med. 2011 Oct26;3(106):106ra106.
- Ravel J, Gajer P, Abdo Z, Schneider GM, Koenig SS, McCulle SL, Karlebach S, Gorle R, Russell J, Tacket CO, Brotman RM, Davis CC, Ault K, Peralta L, ForneyLJ. Vaginal microbiome of reproductive-age women. Proc Natl Acad Sci U S A. 2011 Mar 15;108 Suppl 1:4680-7.
- Hummelen R, Fernandes AD, Macklaim JM, Dickson RJ, Changalucha J, Gloor GB, and Reid G. Deep sequencing of the vaginal microbiome in HIV patients. PLoS One 2010 5(8): e12078.
- Ngugi BM, Hemmerling A, Bukusi EA, Kikuvi G, Gikunju J, Shiboski S, Fredricks DN, Cohen CR. Effects of bacterial vaginosis-associated bacteria and sexual intercourse on vaginal colonization with the probiotic Lactobacillus crispatus CTV-05. Sex Transm Dis. 2011 Nov;38(11):1020-7.
- Martinez RC, Franceschini SA, Patta MC, Quintana SM, Gomes BC, De Martinis EC, Reid G. Improved cure of bacterial vaginosis with single dose of tinidazole (2g), Lactobacillus rhamnosus GR-1, and Lactobacillus reuteri RC-14: a randomized, double-blind, placebo-controlled trial. Can J Microbiol. 2009 Feb;55(2):133-8.
- Martinez RC, Franceschini SA, Patta MC, Quintana SM, Candido RC, Ferreira JC, De Martinis EC, Reid G. Improved treatment of vulvovaginal candidiasis with fluconazole plus probiotic Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14. Lett Appl Microbiol. 2009 Mar;48(3):269-74.
- Anukam K, Osazuwa E, Ahonkhai I, Ngwu M, Osemene G, Bruce AW, Reid G. Augmentation of antimicrobial metronidazole therapy of bacterial vaginosis with oral probiotic Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14: randomized, double-blind, placebo controlled trial. Microbes Infect. 2006 8(6):1450-4.
- Anukam KC, Duru MU, Eze CC, Egharevba J, Aiyebelehin A, Bruce AW, Reid G. Oral use of probiotics as an adjunctive therapy to fluconazole in the treatment of yeast vaginitis: a study of Nigerian women in an outdoor clinic. Microbial Ecol Health Dis. 2009 21: 72-77.
- Koehler G, Assefa S, Reid G. Probiotic interference of Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14 with the opportunistic fungal pathogen Candida albicans. Infect. Dis. Obstetr. Gynecol. 2012 636474.
- Donders GG, Vereecken A, Bosmans E, Dekeersmaecker A, Salembier G, Spitz B. Definition of a type of abnormal vaginal flora that is distinct from bacterial vaginosis: aerobic vaginitis. BJOG. 2002 Jan;109(1):34-43.
- Cotar AI, Chifiriuc MC, Dinu S, Pelinescu D, Banu O, Lazar V. Quantitative real-time PCR study of the influence of probiotic culture soluble fraction on the expression of Pseudomonas aeruginosa quorum sensing genes. Roum Arch Microbiol Immunol. 2010 Oct-Dec;69(4):213-23.
- Corr SC, Li Y, Riedel CU, O’Toole PW, Hill C, Gahan CG. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc Natl Acad Sci U S A. 2007 May 1;104(18):7617-21.
- Reid G, Younes JA, Van der Mei HC, Gloor GB, Knight R, Busscher HJ. Microbiota restoration: natural and supplemented recovery of human microbial communities. Nat Rev Microbiol. 2011 Jan;9(1):27-38.
- Li J, Wang W, Xu SX, Magarvey NA, McCormick JK. Lactobacillus reuteri-produced cyclic dipeptides quench agr-mediated expression of toxic shock syndrome toxin-1 in staphylococci. Proc Natl Acad Sci U S A. 2011 Feb22;108(8):3360-5.
- Younes JA, van der Mei HC, van den Heuvel E, Busscher HJ, Reid G. Adhesion forces and coaggregation between vaginal staphylococci and lactobacilli. PLoS One. 2012;7(5):e36917.
- Kirjavainen PK, Laine RM, Carter D, Hammond JA, Reid G. Expression of anti-microbial defense factors in vaginal mucosa following exposure to Lactobacillus rhamnosus GR-1. Int. J. Probiotics 2008 3: 99-106.
- Yeganegi M, Watson CS, Martins A, Kim SO, Reid G, Challis JR, Bocking AD. Effect of Lactobacillus rhamnosus GR-1 supernatant and fetal sex on lipopolysaccharide-induced cytokine and prostaglandin-regulating enzymes in human placental trophoblast cells: Implications for treatment of bacterial vaginosis and prevention of preterm labour. Amer. J. Obstetr. Gynecol. 2009 May;200(5):532.e1-8. Epub 2009 Mar 14.
- Yeganegi, M, Leung CG, Martins A, Kim SO, Reid G, Challis JR, Bocking AD. Lactobacillus rhamnosus GR-1-induced IL-10 production in human placental trophoblast cells involves activation of JAK/STAT and MAPK pathways. Reprod. Sci. 2010 17(11):1043-51.
- Yeganegi, M, Leung CG, Martins A, Kim SO, Reid G, Challis JR, Bocking AD. Lactobacillus rhamnosus GR-1 stimulates colony stimulating factor 3 (granulocyte) (CSF3) output in placental trophoblast cells in a fetal sex-dependant manner. Biol. Reprod. 2011 84(1):18-25.
- Bisanz J, Reid G. Unraveling how probiotic yogurt works. Sci. Trans. Med. 2011 Oct 26;3(106):106ps41.
SKIN MICROBIOTA: A SOURCE OF DISEASE OR DEFENSE?
Dr Jürgen Schauber, Department of Dermatology and Allergy, Ludwig-Maximilian University Munich, Germany.
Our skin is a natural defence barrier and elements important in this function include:
• physical barrier
• chemical barrier
• peptide barrier
• recognition system
• communication system
• resident/non-resident cells
The epidermis is in effect an immune organ of considerable complexity. At the same time the skin has a resident microbial flora which is always present and only occasionally causes problems.
ANTIMICROBIAL PEPTIDES
The skin is exposed to many pathogens including bacteria, viruses and fungi that cause familiar diseases. One element of protection against these is the production of antimicrobial peptides, constituting an innate antimicrobial defense mechanism.
Peptides with antimicrobial activity (AMPs) are found throughout nature in plants, insects, amphibians and mammals. Many proteins and peptides with antimicrobial properties (in vitro) have been found in skin. Table 1 gives a partial listing of these.
Table 1: Many peptides and proteins in the skin have antimicrobial activity (partial listing)

There is evidence that these peptides, although they may have other functions, do indeed have a role in antimicrobial defence. For example, studies in transgenic mice that cannot produce cathelicidins suggest that these peptides are an important native component of innate host defence in mice and provide protection against invasive skin infection caused by Group A Streptococcus (GAS)1. Human keratinocytes secrete the S100 protein psoriasin which appears to have an important role in inhibiting E. coli survival on human skin2.
AMPs are induced in skin in response to injury or infection3. Cells at wound edges express cell surface AMPS which kill bacteria as part of a nonimmune defense mechanism. These AMPs have been demonstrated to have broad antimicrobial activity against a range of potential pathogens. After secretion a single AMP gene product or ‘pro-peptide’ may be processed into multiple peptides with various antimicrobial properties, which may make it more difficult for pathogens to generate resistance4.
Cutaneous AMPs such as the cathelicidins act as signaling molecules (figure 1) to promote a wide variety of immune functions. Thus as well as being inducible elements of innate immunity in skin they not only function as endogenous antibiotics but also as ‘alarmins’ to prime other repair and defence processes.
Figure 1: Cutaneous AMPs such as the cathelicidins act as signaling molecules

Greater understanding of the role of AMPs in cutaneous innate immune function has led to exploration of their role in patients where these functions appear disordered. For example atopic individuals suffer cutaneous inflammation and are susceptible to inflammation leading some workers to question whether expression of AMPs is normal in atopic conditions. Early work in this field supported the hypothesis that there may be decreased expression of AMPs in atopics5. However later work in atopic eczema showed enhanced expression of the human cathelicidin LL-37 in lesional skin compared with nonlesional skin6, leading the authors to suggest that LL-37 might be associated with the process of re-epithelialization. Recent work suggests that injury downregulates the expression of cathelicidin protein hCAP18/LL-37 in atopic dermatitis (AD)7. Since itching is a primary symptom of AD and scratching inevitably injures the skin, failure to upregulate AMPs in eczema following injury is likely to affect antimicrobial protection and tissue repair in AD.
REGULATION OF AMP PRODUCTION IN SKIN
Toll-like receptors (TLRs), thanks in part to the pioneering work of Jules Hoffmann and Bruce Beutler, are now known to play an important role in alerting the immune system to the presence of a variety of molecules associated with microbial pathogens (figure 2).
Figure 2: Toll-like receptors
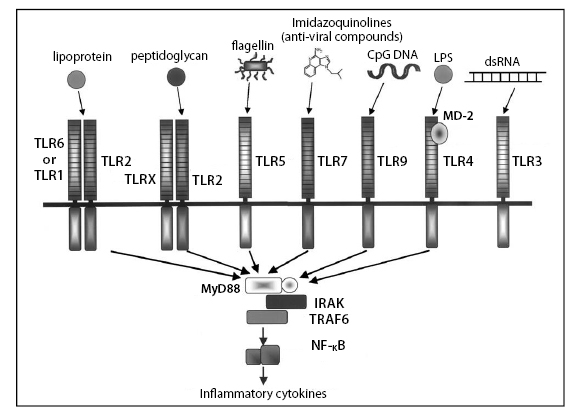
Activation of TLRs induce AMP in skin and mediators of inflammation induce AMP expression (figure 3).
Figure 3:

The search for cathelicidin AMP inducing factors has revealed an important role for dietary sources of cholecalciferol (Vitamin D3)8 including fatty fish species (e.g. catfish, salmon, mackerel, sardines, tuna, eel), whole egg, beef liver and fish liver oils, such as cod liver oil. Some foods, notably milk, dairy products and fruit juice are now also fortified with the vitamin.
The metabolism of Vitamin D3 takes place partly in the skin (figure 4) and this substance is now known to have an important role in some key functions of skin:
• VD3 regulates proliferation, differentiation of keratinoytes
• VD3 analogs have been used for treatment of inflammatory skin diseases
• VD3 effects adaptive immunity
• Keratinocytes produce VD3
• VD3 influences innate immunity in skin
Figure 4:

Vitamin D metabolism may be activated in skin wounds. CYP27B1 – which activates 1,25D3 – is induced in wounds 24 hours after injury and is induced by factors of the wound micromilieu in vitro. Vitamin D3 induces TLR2, CD14 in keratinocytes and enables increased innate immune responses and increased antimicrobial activity. Topical vitamin D induces cathelicidin AMP in human skin and oral vitamin D induces cathelicidin AMP in human skin in atopic patients9. The current model of cathelicidin AMP activation in keratinocytes is illustrated in figure 5.
Figure 5: Current model of cathelicidin AMP activation in keratinocytes
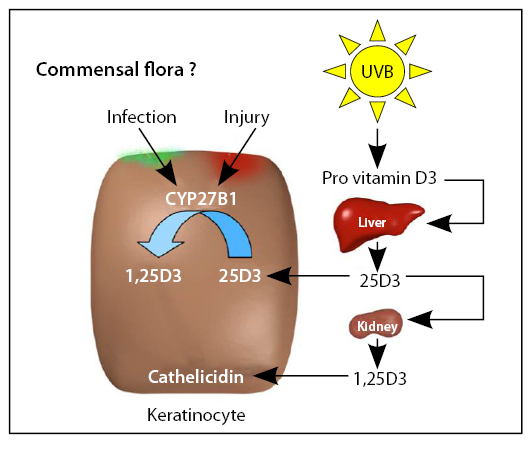
In summary AMPs are regulated by TLR activation, skin inflammation and vitamin D. This opens up some intriguing possibilities for future treatments with possible roles for vitamin D supplementation and TLR activation in the treatment of atopic inflammation and infectious skin disease10.
ROLE OF COMMENSAL FLORA
Recent work shows that skin commensals affect AMP expression by keratinocytes11. Activation of TLR2 by a small molecule produced by S. epidermidis increases antimicrobial defence against bacterial skin infection. Commensals have also been shown to produce AMPs: phenol-soluble modulins (PSM) are secreted by S. epidermidis and selectively kill pathogens such as Streptococci (GAS)12. Skin commensal staphylococci amplify the innate immune response to pathogens by activation of distinct signaling pathways13. Also S. epidermidis-derived delta-toxin cooperates with the host-derived antimicrobial peptides in the innate immune system to reduce survival of pathogenic group A Streptococcus14. AMPs produced by epithelial cells are differentially processed and inactivated by commensals15 (in this study Finegoldia magna) and pathogens (Streptococcus pyogenes).
In summary, commensals such as S. epidermidis:
• produce AMPs to kill pathogens
• produce toxins to enhance AMP activity
• process AMPs to fight pathogens
• activate TLRs to induce cutaneous AMP production
• activate signaling pathways to induce AMPs
So the commensal bacteria resident on the skin can certainly be added to the important elements that constitute the innate immunity of skin.
The work done so far leads to some stimulating open questions including:
Which commensals are the most beneficial ?
What factors of commensals regulate AMPs ?
How can beneficial commensals be supported ?
When we can answer these questions we will be closer to knowing how we can exploit these mechanisms for the prevention and/or therapy of infectious skin disease or skin diseases associated with barrier dysfunction.
References
- Nizet V, Ohtake T, Lauth X, Trowbridge J, Rudisill J, Dorschner RA, Pestonjamasp V, Piraino J, Huttner K, Gallo RL. Innate antimicrobial peptide protects the skin from invasive bacterial infection. Nature. 2001 Nov 22;414(6862):454-7.
- Gläser R, Harder J, Lange H, Bartels J, Christophers E, Schröder JM. Antimicrobial psoriasin (S100A7) protects human skin from Escherichia coli infection. Nat Immunol. 2005 Jan;6(1):57-64.
- Gallo RL, Ono M, Povsic T, Page C, Eriksson E, Klagsbrun M, Bernfield M. Syndecans, cell surface heparan sulfate proteoglycans, are induced by a proline-rich antimicrobial peptide from wounds. Proc Natl Acad Sci U S A. 1994 Nov 8;91(23):11035-9.
- Murakami M, Lopez-Garcia B, Braff M, Dorschner RA, Gallo RL. Postsecretory processing generates multiple cathelicidins for enhanced topical antimicrobial defense. J Immunol. 2004 Mar 1;172(5):3070-7.
- Ong PY, Ohtake T, Brandt C, Strickland I, Boguniewicz M, Ganz T, Gallo RL, Leung DY. Endogenous antimicrobial peptides and skin infections in atopic dermatitis. N Engl J Med. 2002 Oct 10;347(15):1151-60.
- Ballardini N, Johansson C, Lilja G, Lindh M, Linde Y, Scheynius A, Agerberth B. Enhanced expression of the antimicrobial peptide LL-37 in lesional skin of adults with atopic eczema. Br J Dermatol. 2009 Jul;161(1):40-7. Epub 2009 Mar 20.
- Mallbris L, Carlén L, Wei T, Heilborn J, Nilsson MF, Granath F, Ståhle M. Injury downregulates the expression of the human cathelicidin protein hCAP18/LL-37 in atopic dermatitis. Exp Dermatol. 2010 May;19(5):442-9. Epub 2009 Jul 23.
- Wang TT, Nestel FP, Bourdeau V, Nagai Y, Wang Q, Liao J, Tavera-Mendoza L, Lin R, Hanrahan JW, Mader S, White JH. Cutting edge: 1,25-dihydroxyvitamin D3 is a direct inducer of antimicrobial peptide gene expression. J Immunol. 2004 Sep1;173(5):2909-12. Erratum in: J Immunol. 2004 Nov 15;173(10):following 6489. Hanrahan, JH [corrected to Hanrahan, JW].
- Hata TR, Kotol P, Jackson M, Nguyen M, Paik A, Udall D, Kanada K, Yamasaki K, Alexandrescu D, Gallo RL. Administration of oral vitamin D induces cathelicidin production in atopic individuals. J Allergy Clin Immunol. 2008 Oct;122(4):829-31.
- Gombart AF. The vitamin D-antimicrobial peptide pathway and its role in protection against infection. Future Microbiol. 2009 Nov;4(9):1151-65. Review.
- Lai Y, Cogen AL, Radek KA, Park HJ, Macleod DT, Leichtle A, Ryan AF, Di Nardo A, Gallo RL. Activation of TLR2 by a small molecule produced by Staphylococcus epidermidis increases antimicrobial defense against bacterial skin infections. J Invest Dermatol. 2010 Sep;130(9):2211-21. Epub 2010 May 13.
- Cogen AL, Yamasaki K, Sanchez KM, Dorschner RA, Lai Y, MacLeod DT, Torpey JW, Otto M, Nizet V, Kim JE, Gallo RL. Selective antimicrobial action is provided by phenol-soluble modulins derived from Staphylococcus epidermidis, a normal resident of the skin. J Invest Dermatol. 2010 Jan;130(1):192-200. Epub .
- Wanke I, Steffen H, Christ C, Krismer B, Götz F, Peschel A, Schaller M, Schittek B. Skin commensals amplify the innate immune response to pathogens by activation of distinct signaling pathways. J Invest Dermatol. 2011 Feb;131(2):382-90. Epub 2010 Nov 4.
- Cogen AL, Yamasaki K, Muto J, Sanchez KM, Crotty Alexander L, Tanios J, Lai Y, Kim JE, Nizet V, Gallo RL. Staphylococcus epidermidis antimicrobial delta-toxin (phenol-soluble modulin-gamma) cooperates with host antimicrobial peptides to kill group A Streptococcus. PLoS One. 2010 Jan 5;5(1):e8557.
- Frick IM, Nordin SL, Baumgarten M, Mörgelin M, Sørensen OE, Olin AI, Egesten A. Constitutive and inflammation-dependent antimicrobial peptides produced by epithelium are differentially processed and inactivated by the commensal Finegoldia magna and the pathogen Streptococcus pyogenes. J Immunol. 2011 Oct15;187(8):4300-9. Epub 2011 Sep 14.
THE SKIN - MICROBES AND INFECTION
Professor Roderick J. Hay, Kings College Hospital, London and International Foundation for Dermatology, UK.
THE SKIN MICROBIOTA
The human skin is a normal habitat for many microbes including bacteria – Staph epidermidis, Propionibacterium acnes, micrococci, coryneforms, brevibacteria, and fungi – e.g. Malassezia species, but no viruses. Among ectoparasites, only Demodex folliculorum is a frequent resident.
As well as the ‘resident’ microbiota, normal skin may be a temporary site of carriage for other bacteria and fungi e.g. Staph aureus, Candida albicans, and intermittently occurring residence, as with the Herpes simplex virus. The presence of diseased or damaged skin (e.g. in eczema) may also change the nature of the resident microbiota.
The commensal microbiota of the skin can, on occasions, cause problems. For example contamination of transdermal medical procedures e.g. with Staph epidermidis can be a source of septicaemia in neutropenic patients. In other cases, with e.g. Propionibacterium acnes, and Malassezia, alteration in local conditions in the skin changes the way the microbe behaves in ways that can cause problems for the host.
The skin has a natural defence system and factors including epidermal growth, antimicrobial peptides (e.g. cecropins and magainins), fatty acids, pH, and immunity (both epidermal/systemic) contribute to the barrier against pathogens.
Skin infections are common and can cause considerable morbidity. Table 1 shows the global burden of illness and cost per disability-adjusted life year (DALY) of treating the three commonest skin diseases caused by infection.
Table 1: Cost of Cure and Impact on DALY’s- 3 commonest diseases

SKIN INFECTIONS
There are 4 stages common to all microbial skin infections illustrated in figure 1, but examining specific infections is instructive.
Figure 1: Pathogenesis of microbial infection of skin

MALASSEZIA
Malassezia is a commensal yeast on skin – particularly on hair-bearing skin of the scalp and upper trunk. It is a mainly lipophilic microbe with a characteristic cell wall structure and polar bud formation and has been known to causes disease in animals and humans. It is associated with specific enzymes e.g. phospholipases and melanin formation.
The complete Malassezia globosa and M restricta genomes have now been sequenced. That of M. globosa represents the smallest genomic of a free-living eukaryotic cell – 300 times smaller than the human genome. The closest relative to this organism is Ustilago maydis – common crop pathogen (corn smut).
Malassezia spp (including M.globosa, sympodialis, furfur, restricta, slooffiae, obtusa, dermatitidis, nana) have been associated with a number of skin diseases:
• Pityriasis versicolor
• Malassezia folliculitis
• Seborrhoeic dermatitis
• Atopic dermatitis affecting the head and neck
• Deep infection (only in neonatal period)
PITYRIASIS VERSICOLOR
This common and harmless skin condition, characterised by a fine scaly rash that gives the skin a variable colour (pink or fawn on pale skin but with pale patches on tanned skin) is usually caused by M. globosa. Spontaneous remission is rare. There is no evidence of activation of T lymphocyte responses in Pityriasis versicolor, suggesting ineffective immunity. Suggested reasons for this include diversion of immune cells, and importantly, inhibition of immune systems by yeast lipid.
SEBORRHOEIC DERMATITIS AND DANDRUFF
The role of Malassezia spp in seborrhoeic dermatitis and dandruff has an interesting history. In 1874 Malassez described a fungus associated with dandruff or pilade. In 1976 Leyden and Kligman conducted some experiments and concluded: ‘The studies demonstrate that the increased number of scalp microorganisms found in dandruff occurs as a secondary event to increased nutrients and that scalp organisms play no primary role in the pathogenesis of dandruff’1. In 1984, Shuster essentially ‘re-asserted’ the central role of this fungus in dandruff, upon which current antifungal treatments for this condition is based2.
Seborrhoeic dermatitis and dandruff have typical clinical features of erythema, greasy scales and a typical distribution – involving scalp, eyebrows, behind the ears, and front of the chest. Seborrhoeic dermatitis is a severe form of dandruff but involving other skin sites.
Seborrhoeic dermatitis (SD) is a common condition but also has a number of clinical associations with more serious disease – including tertiary syphilis, chronic neurological disease and AIDS. There appears to be no correlation between species and disease i.e. allMalassezia spp have been grown from SD, but three dominate: M. globosa (only certain genetic strains), M. furfur and M. restricta.
In immunological studies no correlation is seen between T or B lymphocyte activity or cell infiltrates and disease. In SD patients, unaffected and affected skin show similar cytokine and immune cells profiles (IL-2 and IL-6 to IFN γ and TNF α). Increased numbers of organisms are observed in AIDS and this is associated with the CD4 count but not the presence of SD.
Sebum has been thought to play a role in dandruff and SD, but sebum composition in patients is highly variable and unrelated to Malassezia colonisation or disease severity. However M. globosa can produce oleic acid on dandruff-susceptible individuals.
The results of genomics and proteomics show that the M. globosa genome encodes for 14 lipases, and 9 phospholipases. Most lipases are expressed on the scalp, but not in culture medium and 8 lipases and 3 phospholipases have been identified. We can conclude that secreted enzymes probably enable survival on the scalp but may damage scalp cells.
Malassezia behaves differently in normal and SD skin with higher production of metabolites with immunological activity such as malassezin, and indolcarbazol from M. furfur isolates in SD compared to normal skin3.
Figure 2 provides a summary of the known immunological interactions in pityriasis versicolor and seborrhoeic dermatitis.
Figure 2: Summary of immunological events in pityriasis versicolor and seborrhoeic dermatitis

BACTERIAL INFECTIONS OF THE SKIN
Bacteria can cause skin infections of varying severity and the following is a non-exhaustive list:
• Impetigo – Staph aureus, streptococci
• Folliculitis/furunculosis ( boils )
• Cellulitis and erysipelas
• Necrotizing fasciitis
• Erythrasma
• Gram negative infections
• Cutaneous Tuberculosis
• Rarities eg bacillary angiomatosis
• Secondary bacterial infection eg eczema
STAPHYLOCOCCAL SKIN INFECTION
Impetigo, boils, and folliculitis are caused by Staphylococcus aureus (infections can sometimes cause staphylococcal scalded skin syndrome (SSSS) and toxic shock associated with the production of exotoxins). In 1985 in the UK, 90% of S. aureus were resistant to penicillin (ß- lactam) 32% to erythromycin, and 76% to tetracycline.
Multiple drug resistance in staphylococci is a major and growing problem (figure 3)4 and is now subdivided into:
• Hospital acquired methicillin resistant Staphylococcus aureus – HA-MRSA
• Community acquired methicillin resistant Staphylococcus aureus – CA-MRSA
Figure 3: Spread of MRSA associated with skin and soft tissue infection (SSTI) 2001-05 Baltimore
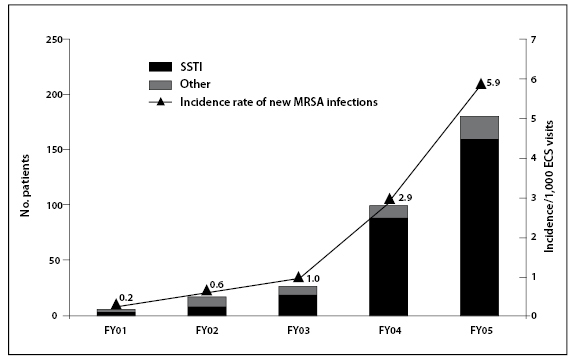
Reproduced with permission from Emerging Infectious Diseases. Johnson et al. 2007 August; 13(8): 1195–1200.
Table 2 summarises the difference between HA and CA MHRSA. The latter has a particular propensity to cause skin and soft tissue infections and, in contrast to HA-MRSA, occurs often in immunocompetent individuals.
Table 2: Differences between Hospital-acquired ( HA ) versus Community-acquired (CA ) methicillin resistant Staph aureus ( MRSA )
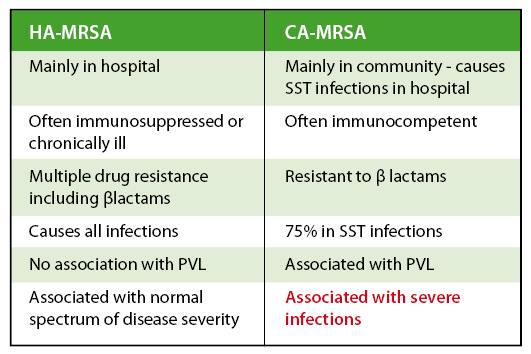
Panton Valentine Leukocidin (PVL) in sensitive strains has been identified as a risk factor for severe infection. This cytotoxin causes experimental skin necrosis in animals. It works through synergy between 2 proteins LukS-PV and LukF-PV and may be present even if the bacterial strains are sensitive to antibiotics. Treatment is often ineffective, and the presence of PVL strains may only be recognised by a clinical pattern of poor response, frequent relapse, and severe lesions, often in otherwise healthy patients. There seems to be an association with recent travel.
Del Giudice et al5 reported a series of Staph aureus cases from a dermatology department in France during 2003-2010. They found that 74% of follicular infections compared to 12% of non-follicular infections were caused by PVL positive strains. In a survey of skin and soft tissue infections (SSTI) from two General Practice regions in the Netherlands Mithoe et al6found that 11% of patients with Staph aureus were PVL positive. PVL strains were commoner in the North than the South region, perhaps reflecting regional differences e.g. in antibiotic usage.
Practical steps to take in dealing with this problem include:
• Suspect early if relapse is frequent and there are extensive and/or severe lesions.
• Have second line antibiotics available – e.g. rifampicin, clindamycin plus antisepsis e.g. povidone or hibisol.
• Warn the local microbiology laboratory if there is a suspicion of PVL.
ATOPIC DERMATITIS
Atopic dermatitis (AD) is common in most countries with a prevalence in 11-13 year olds approaching 12-18%. There are theories that early exposure to antigens may play a protective role in development of the condition. Over 80% of AD shows colonisation withStaph aureus. Observation of improvement on antibiotics correlating with reduction of flares raises the question ‘is this infection’? Mechanisms for a role of infection in AD include raised anti-Staph IgE, reduced antimicrobial peptide production and the presence of ‘superantigens’ e.g. α toxin, peptidoglycan7 and lipoteichoic acid.
ECTOPARASITES AND THE SKIN
Demodex is a genus of microscopic parasitic mites that live in or near the hair follicles of mammals. The role of these ectoparasites in human diseases is disputed but there are reports of clinical cases involving Demodex in HIV and immunosuppressed patients. Other occasional human ectoparasites resident in or on the body include scabies and head lice.Cheyletiella, from cats/dogs can survive for hours on human skin, and others e.g. bed bugs and mosquitos, do not live on skin, but visit only to feed. The importance of some of these parasites lies in their role as potential vectors for transmission of disease.
PYODERMA, SCABIES AND RENAL DAMAGE
There is an association between scabies and Group A streptococcal infection in the tropics.In outbreaks of scabies associated with pyoderma, symptomatic nephritis or haematuria/proteinuria can occur in 5-10% of cases. 10 years later 8-21% of affected children still have haematuria and previous scabies is associated with raised urinary albumin : creatinine ratio. This concurrence of infestation and infection has also been associated with rheumatic fever and infant septicaemia.
SUMMARY OF RESEARCH TARGETS FOR THE FUTURE
The challenges of skin infection and infestation remain considerable. As we understand better the mechanisms involved in maintaining the normal microbiota and repelling pathogens, interesting possibilities will undoubtedly emerge.
Potential research targets, including targets for manipulation of microbial flora of the skin and/or probiotics in the future might include:
• Recolonising to prevent infection eg Staph aureus, Malassezia – complete replacement or reduction.
• Exploring the relationship between microbiota elsewhere and skin disease e.g. chronic urticaria/Helicobacter
• Other areas unrelated to infection e.g. generation of key nutrients e.g. Vitamin D
• Can one replace and /or use bacteria in other ways e.g. prevention of insect vector biting, changing endozootic bacteria to affect the viability of the ectoparasite.
References
- Leyden JJ, McGinley KJ, Kligman AM. Role of microorganisms in dandruff. Arch Dermatol. 1976 Mar;112(3):333-8.
- Shuster S. The aetiology of dandruff and the mode of action of therapeutic agents. Br J Dermatol. 1984 Aug;111(2):235-42. Review.
- Gaitanis G, Magiatis P, Stathopoulou K, Bassukas ID, Alexopoulos EC, Velegraki A, Skaltsounis AL. AhR ligands, malassezin, and indolo[3,2-b]carbazole are selectively produced by Malassezia furfur strains isolated from seborrheic dermatitis. J Invest Dermatol. 2008 Jul;128(7):1620-5. Epub 2008 Jan 24.
- Johnson JK, Khoie T, Shurland S, Kreisel K, Stine OC, Roghmann MC. Skin and soft tissue infections caused by methicillin-resistant Staphylococcus aureus USA300 clone. Emerg Infect Dis. 2007 Aug;13(8):1195-200.
- Del Giudice P, Bes M, Hubiche T, Blanc V, Roudière L, Lina G, Vandenesch F, Etienne J. Panton-Valentine leukocidin-positive Staphylococcus aureus strains are associated with follicular skin infections. Dermatology. 2011;222(2):167-70. Epub 2011 Feb 22.
- Mithoe D, Rijnders MI, Roede BM, Stobberingh E, Möller AV. Prevalence of community-associated meticillin-resistant Staphylococcus aureus and Panton-Valentine leucocidin-positive S. aureus in general practice patients with skin and soft tissue infections in the northern and southern regions of The Netherlands. Eur J Clin Microbiol Infect Dis. 2012 Mar;31(3):349-56. Epub 2011 Jun 18.
- Matsui K, Nishikawa A. Percutaneous application of peptidoglycan from Staphylococcus aureus induces infiltration of CCR4+ cells into mouse skin. J Investig Allergol Clin Immunol. 2011;21(5):354-62.
LACTOBACILLI AND PROBIOTICS IN GYNECOLOGY AND OBSTETRICS
Professor Werner Mendling, Vivantes – Klinikum im Friedrichshain and Am Urban, Clinics for Obstetrics and Gynecology, Berlin, Germany.
Of the approximately 120 Lactobacillus species known, about 15 are known to occur in the vagina. These bacteria act on glycogen in the vaginal epithelium to produce lactic acid which helps to keep the pH in the vagina in the range of 3.8-4.4 (although some women with nolactobacilli have normal vaginal pH)*. Lactobacilli are now known to produce other substances besides lactic acid including:
• Hydrogen peroxide (H2O2)
• bacteriocins
• biosurfactants
• coaggregation molecules
• and other as yet unknown substances
As well as Lactobacillus species, the normal vaginal flora can include transient species including: Gardnerella vaginalis, Atopobium vaginae, Mycoplasmas Bacteroides-, Prevotella– and Porphyromonas species, Peptostreptococci, and Candida albicans among many others.
Bacterial vaginosis is the most common cause of abnormal vaginal discharge in women of childbearing age. This syndrome of unknown cause is characterized by depletion of the normal Lactobacillus population and an overgrowth of vaginal anaerobes, accompanied by loss of the usual vaginal acidity. Women with symptomatic bacterial vaginosis report an offensive, fishy-smelling discharge that is often most noticeable after unprotected intercourse or at the time of menstruation.
When bacterial vaginosis develops the pH rises above 4.5, to levels up to 6.0. The lactobacilli reduce in concentration and may disappear, while there is an increased concentration of anaerobic and facultative anaerobic organisms. This pattern is illustrated in table 1.
Table 1:
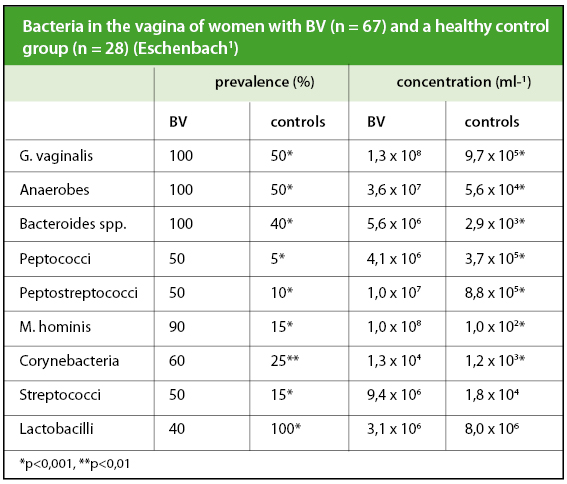
There are several species of Lactobacillus present in normal women and the most important include L. crispatus, L. gasseri, L. jensenii, and L. iners but not L. acidophilus.
Diagnosis of BV is confirmed by identifying at least 3 of 4 ‘Amsel criteria’:
• The typical thin homogenous grey vaginal discharge
• Vaginal pH > 4.5
• Clue cells detected on microscopy of vaginal fluid. Epithelial cells covered with so many small bacteria that the border is fuzzy are termed ‘clue cells’, because their presence is a clue to the diagnosis.
• Release of a strong fishy odour when vaginal fluid is mixed with alkali (usually 10% potassium hydroxide) – the whiff test.
Generally in research studies the Nugent scoring system is used to interpret Gram-stained vaginal smears. A score of between 0 and 10 is generated from counting bacterial morphotypes with 0-3 being normal, 4-6 intermediate and 7-10 indicating BV.
The prevalence of BV varies markedly by population and geography:
Germany, pregnant women 20 %
Black women in USA > 30 %
Women in central Africa > 50 %
Although many women with BV are asymptomatic, presence of BV markedly increases the relative risk of gynaecological infection (pelvic inflammatory disease and infection after hysterectomy) and obstetric complications (pre-term birth and post-partum endometritis). Several studies2,3 have confirmed that treatment of BV significantly reduces the rate of pre-term birth and late miscarriages.
BIOFILMS IN BV
In a paper published in 20054 we investigated the composition and spatial organization of bacteria associated with the vaginal epithelium in biopsy specimens from 20 patients with bacterial vaginosis and 40 normal premenopausal and postmenopausal controls using a broad range of fluorescent bacterial group-specific rRNA-targeted oligonucleotide probes. Bacterial vaginosis was associated with greater occurrence and higher concentrations of a variety of bacterial groups. However, only Gardnerella vaginalis together with Atopobium vaginae developed a characteristic adherent biofilm that was specific for bacterial vaginosis (figure 1).
Figure 1: Gardnerella vaginalis biofilm

We concluded that a biofilm comprised of confluent G. vaginalis with other bacterial groups incorporated in the adherent layer is a prominent feature of bacterial vaginosis.
Subsequently we showed that an adherent Gardnerella vaginalis biofilm persists on the vaginal epithelium following standard therapy of bacterial vaginosis with oral metronidazole5. This seems to serve as a reservoir of the core pathogens for BV. Although the biofilm primarily consisted of Gardnerella vaginalis and Atopobium vaginae, Lactobacilli andBacterioides/Prevotella (7/16) were also present but accounted for only < 10 % of all bacteria.
BV therapeutic options include:
• antibiotics (metronidazole, clindamycin)
• lactic acid
• lactobacilli
• Dis-infective agents
• Preventative lactobacillus vaccination
Numerous products aimed at Lactobacillus substitution exist on the market in Germany. Well investigated strains for vaginal or oral treatment include L. rhamnosus GR-1 and L. reuteri RC-14 (formerly L. fermentum) or L. rhamnosus GG and others.
VAGINAL ADMINISTRATION
A study of L. rhamnosus GG, administered vaginally, in 10 subjects, showed that the bacteria survived only 5 days in situ, but L. rhamnosus GR-1 and L. reuteri RC-14 were detectable for 19 days6. This highlights the importance of selection of strains for urogenital probiotic applications.
A double-blind, randomized, placebo-controlled study of vaginal probiotics for recurrent BV7investigated the administration of L. rhamnosus, L. acidophilus, Strept. thermoph. for 7 days on, 7 days off, and then 7 days on. The study showed a significant reduction of recurrences of BV in the active vs placebo group (15.8 vs. 45.0 %) and reduction of G. vaginalisincidence (3.5 vs. 18.3 %) after 2 and 11 months.
A phase 2a study8 assessed colonization efficiency, safety, tolerability, and acceptability ofLactobacillus crispatus CTV-05 administered by a vaginal applicator. Twenty-four women with BV were randomized in a 3:1 ratio of active product to placebo. Participants usedLactobacillus crispatus CTV-05 at 102 colony-forming units (cfu)/dose or placebo for 5 initial consecutive days, followed by a weekly application over 2 weeks. Sixty-one percent of the 18 women randomized to the active group were colonized with L. crispatus CTV-05 at Day 10 or Day 28. The treatment was well tolerated and accepted.
Antonio et al.9 investigated two potencies of gelatin capsules containing Lactobacillus crispatus CTV-05 for safety and vaginal colonization in 90 young women. Of 40 participants who lacked L. crispatus colonization at enrollment, 36 (90%) were successfully colonized by CTV-05 at 1 or more follow-up visits, whereas only 24 (51%) of 47 participants colonized byL. crispatus at enrollment were positive for CTV-05 at follow-up (P < .001). The authors concluded that that the factors that predict failure to become colonized by probiotic lactobacilli include exposure to semen, vaginal intercourse, and the presence of lactobacilli of the same species at enrollment.
Supplemental vaginal treatment with two vaginal lactobacilli strains for 3 months did not improve clindamycin therapy of BV, but reduced the relapse rate of initially cured women significantly after 6 months10. A double-blind, placebo-controlled study11, showed that use of tampons with L. fermentum, L. casei var. rhamnosus and L. gasseri for 4 weeks after vaginal clindamycin in BV patients did not improve therapy results of BV after two months, but improved significantly results of Nugent-Score 4 – 6 patients.
In a study in Nigeria12, 40 women diagnosed with BV were randomized to receive either two dried capsules containing Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14each night for 5 days, or 0.75% metronidazole gel, applied vaginally twice a day (in the morning and evening). Follow-up at day 6, 15 and 30 showed cure of BV in significantly more probiotic treated subjects (16, 17 and 18/20, respectively) compared to metronidazole treatment (9, 9 and 11/20: P=0.016 at day 6, P=0.002 at day 15 and P=0.056 at day 30).
ORAL TREATMENT
In a study by Reid et al.14, forty-two healthy women were randomized to receive one of three encapsulated Lactobacillus rhamnosus GR-1 plus Lactobacillus fermentum RC-14 probiotic dosage regimens or L. rhamnosus GG by mouth each day for 28 days. However, the vaginal flora, assessed by Nugent scoring, was only normal in 40% of the cases, and 14 patients had asymptomatic bacterial vaginosis. Treatment with L. rhamnosus GR-1/L. fermentum RC-14 once and twice daily correlated with a healthy vaginal flora in up to 90% of patients, and 7/11 patients with bacterial vaginosis converted to normal or intermediate scores within 1 month. Ingestion of L. rhamnosus GG failed to have an effect. This study confirms the potential efficacy of orally administered lactobacilli and showed that over 108 viable organisms per day is the required dose.
Another study by the same group15 explored the efficacy of oral lactobacilli. A randomized, placebo-controlled trial of 64 healthy women given daily oral capsules of Lactobacillus rhamnosus GR-1 and Lactobacillus fermentum RC-14 for 60 days showed restoration from asymptomatic bacterial vaginosis microflora to a normal lactobacilli colonized microflora in 37% women during lactobacilli treatment compared to 13% on placebo (P=0.02). Lactobacilli were detected in more women in the lactobacilli-treated group than in the placebo group at 28 day (P=0.08) and 60 day (P=0.05). Culture findings confirmed a significant increase in vaginal lactobacilli at day 28 and 60, a significant depletion in yeast at day 28 and a significant reduction in coliforms at day 28, 60 and 90 for lactobacilli-treated subjects versus controls.
A study from Vienna16 examined the efficacy of orally administered probiotic strainsLactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14 on the quality of the vaginal flora in postmenopausal women in a randomized, double-blind, placebo-controlled study. Twenty-one of the 35 subjects (60%) in the intervention group and 6 of the 37 subjects (16%) in the control group showed a reduction in the Nugent score by at least two grades (p=0.0001). The median difference in Nugent scores between baseline and the end of the study was 3 in the intervention group and 0 in the control group (p=0.0001).
SAFETY
In a review of the safety probiotics, Boyle et al.17 concluded that probiotics have an excellent overall safety record but that they should be used with caution in special risk groups such as patients with immunosuppression or in very small preterm babies with short bowel-syndrome.
Sepsis with oral lactobacilli appears to be extremely rare with n = 7 cases in 1999-2005 withL. rhamnosus, mostly – GG.
* Recent data show significant differences in the predominance of different lactobacillus species in different ethnic groups with significantly different vaginal pH values up to 513.
References
- Eschenbach DA, Davick PR, Williams BL, Klebanoff SJ, Young-Smith K, Critchlow CM, Holmes KK. Prevalence of hydrogen peroxide-producing Lactobacillus species in normal women and women with bacterial vaginosis. J Clin Microbiol. 1989 Feb;27(2):251-6.
- Hoyme UB, Saling E. [Efficient prevention of prematurity - the Thuringian model]. Gynakol Geburtshilfliche Rundsch. 2004 Jan;44(1):2-9. German.
- Kiss H, Petricevic L, Husslein P. Prospective randomised controlled trial of an infection screening programme to reduce the rate of preterm delivery. BMJ.14;329(7462):371. Epub 2004 Aug 4.
- Swidsinski A, Mendling W, Loening-Baucke V, Ladhoff A, Swidsinski S, Hale LP, Lochs H. Adherent biofilms in bacterial vaginosis. Obstet Gynecol. 2005 Nov;106(5 Pt 1):1013-23.
- Swidsinski A, Mendling W, Loening-Baucke V, Swidsinski S, Dörffel Y, Scholze J, Lochs H, Verstraelen H. An adherent Gardnerella vaginalis biofilm persists on the vaginal epithelium after standard therapy with oral metronidazole. Am J Obstet Gynecol. 2008 Jan;198(1):97.e1-6. Epub 2007 Nov 19.
- Gardiner GE, Heinemann C, Bruce AW, Beuerman D, Reid G. Persistence of Lactobacillus fermentum RC-14 and Lactobacillus rhamnosus GR-1 but not L. rhamnosus GG in the human vagina as demonstrated by randomly amplified polymorphic DNA. Clin Diagn Lab Immunol. 2002 Jan;9(1):92-6.
- Ya W, Reifer C, Miller LE. Efficacy of vaginal probiotic capsules for recurrent bacterial vaginosis: a double-blind, randomized, placebo-controlled study. Am J Obstet Gynecol. 2010 Aug;203(2):120.e1-6.
- Hemmerling A, Harrison W, Schroeder A, Park J, Korn A, Shiboski S, Foster-Rosales A, Cohen CR. Phase 2a study assessing colonization efficiency, safety, and acceptability of Lactobacillus crispatus CTV-05 in women with bacterial vaginosis. Sex Transm Dis. 2010 Dec;37(12):745-50.
- Antonio MA, Meyn LA, Murray PJ, Busse B, Hillier SL. Vaginal colonization by probiotic Lactobacillus crispatus CTV-05 is decreased by sexual activity and endogenous Lactobacilli. J Infect Dis. 2009 May 15;199(10):1506-13.
- Larsson PG, Stray-Pedersen B, Ryttig KR, Larsen S. Human lactobacilli as supplementation of clindamycin to patients with bacterial vaginosis reduce the recurrence rate; a 6-month, double-blind, randomized, placebo-controlled study. BMC Womens Health. 2008 Jan 15;8:3.
- Eriksson K, Carlsson B, Forsum U, Larsson PG. A double-blind treatment study of bacterial vaginosis with normal vaginal lactobacilli after an open treatment with vaginal clindamycin ovules. Acta Derm Venereol. 2005;85(1):42-6.
- Anukam KC, Osazuwa E, Osemene GI, Ehigiagbe F, Bruce AW, Reid G. Clinical study comparing probiotic Lactobacillus GR-1 and RC-14 with metronidazole vaginal gel to treat symptomatic bacterial vaginosis. Microbes Infect. 2006 Oct;8(12-13):2772-6. Epub 2006 Sep 11.
- Ravel J, Gajer P, Abdo Z, Schneider GM, Koenig SS, McCulle SL, Karlebach S, Gorle R, Russell J, Tacket CO, Brotman RM, Davis CC, Ault K, Peralta L, Forney LJ. Vaginal microbiome of reproductive-age women. Proc Natl Acad Sci U S A 2011 Mar 15;108 Suppl 1:4680-7. Epub 2010 Jun 3.
- Reid G, Beuerman D, Heinemann C, Bruce AW. Probiotic Lactobacillus dose required to restore and maintain a normal vaginal flora. FEMS Immunol Med Microbiol. 2001 Dec;32(1):37-41.
- Reid G, Charbonneau D, Erb J, Kochanowski B, Beuerman D, Poehner R, Bruce AW. Oral use of Lactobacillus rhamnosus GR-1 and L. fermentum RC-14 significantly alters vaginal flora: randomized, placebo-controlled trial in 64 healthy women. FEMS Immunol Med Microbiol 2003 Mar 20;35(2):131-4
- Petricevic L, Unger FM, Viernstein H, Kiss H. Randomized, double-blind, placebo-controlled study of oral lactobacilli to improve the vaginal flora of postmenopausal women. Eur J Obstet Gynecol Reprod Biol. 2008 Nov;141(1):54-7. Epub 2008 Aug 12.
- Boyle RJ, Robins-Browne RM, Tang ML. Probiotic use in clinical practice: what are the risks? Am J Clin Nutr. 2006 Jun;83(6):1256-64; quiz 1446-7. Review.
LACTOBACILLI IN MUCOSAL CANDIDA ALBICANS INFECTIONS
Professor Martin Schaller Department of Dermatology, Eberhard Karls University Tübingen, Germany.
CANDIDA ALBICANS
Candida albicans is facultative pathogenic yeast which is a fequent colonizer of human skin and mucous membranes. It is present on mucosal surfaces in 40-80% of normal human beings and is part of the microbiota in the mouth, gut, and vagina. This organism is normally a harmless colonizer and infections with Candida usually occur only when a patient has some alteration in cellular immunity, normal flora or normal physiology. Nevertheless Candida albicans is the most common fungal pathogen worldwide and systemic candidosis is the 4th leading cause of nosocomial infections and accounts for 40% of the mortality.
Vulvovaginal candidosis (VVC) is very common and affects 75% of women at some time, with 5% experiencing recurrence. Recurrent VVC, defined as at least four episodes per year, often requires long-term antifungal therapy. In contrast, oral candidosis is most commonly manifested in patients with leukemia or other immunosuppressed conditions and is often a clue to acute primary HIV infection.
Candida albicans has become a cause for concern because it is increasingly resistant to drug therapies. It is also the source of considerable health economic cost, with diagnosis and treatment of VVC and the concomitant loss of productivity estimated to have cost US$1.8 billion in 1995.
What turns Candida albicans from a commensal to a pathogen is a combination of increasing virulence factors in the organism and the interaction of these with predisposing host factors that compromise immunity (figure 1).
Figure 1.

To become a pathogen, the normal commensal Candida albicans must adhere to mucosal surfaces, cause damage to superficial cells and then invade the deeper cell layers (figure 2).
Figure 2: Stages of Mucosal Candidosis
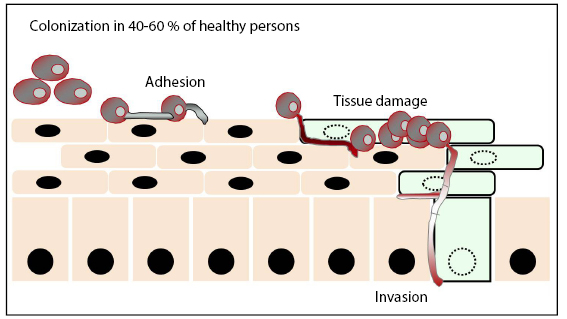
Keratinocytes on mucosal surfaces are the first cells that are in contact with C. albicans and are also the first cells that get damaged by C. albicans. These cells are part of the innate mucosal immune system.
Reconstituted Human Epithelium (RHE) is a valuable research model for the study of Candidal infections1. Infection experiments using these models are highly reproducible and can be used for the direct analysis of pathogen-epithelial cell interactions. To study the impact of innate immunity or the antifungal activity of natural and non-natural compounds, the mucosal infection models can be supplemented with immune cells, antimicrobial agents or probiotic bacteria.
Probiotic Lactobacillus spp. bacteria are widely used in the treatment of vaginal candidiasis. These organisms are suggested to have a number of effects that maintain or regenerate balance between tolerance and defense, including:
• Up-regulation of mucus production
• Improvement of epithelial barrier function
• Increase in IgA production
• Competition for adhesion sites
• Production of organic acids, ammonia, H2O2 and bacteriocins
• Up-regulation of antimicrobial peptides
EFFECTS OF PROBIOTICS ON C. ALBICANS
Lactobacillus rhamnosus GG (LGG) is one of the most important and most investigated probiotic strains and shows protective effects against C. albicans infections2.
Vaginal Lactobacillus-strains have been shown to inhibit the viability of Prevotella bivia andGardnerella vaginalis in cell culture and co-culture experiments3. The mechanism of this inhibition is not attributable to low pH and lactic acid alone, but to hydrogen peroxide and proteolytic enzyme-resistant compound(s) present in the cell-free culture supernatant.
Oral supplementation with L. casei ssp. rhamnosus has also been shown to prevent enteric colonization by Candida species in preterm neonates4. Enhanced clearance of Candida albicans from the oral cavities of mice has been demonstrated following oral administration of Lactobacillus acidophilus5.
In our research programme we set out to answer the following questions:
• Do Lactobacillus spp. protect against mucosal Candida albicans infection?
• What are the mechanisms of protection?
• Is the immune response of the keratinocytes influenced by the probiotic bacteria?
• Is there interaction of probiotic bacteria with virulence of factors of the pathogen?
In culture experiments, we showed only slight inhibition of C. albicans growth by L. rhamnosus GG (LGG) after 72hrs. We also observed that LGG does not block hyphal induction of C. albicans. However in Reconstituted Human Epithelium (RHE) experiments when L. rhamnosus GG is added (at 12-24hrs) to an infection of C. albicans established for 24 hours – a therapeutic model – L. rhamnosus GG mediates protection against C. albicans.
However LGG reduces the mucosal immune response. There are two possible explanations for this, either LGG down-regulates inflammatory response of the keratinocytes or reduced cytokine levels occur as result of reduced cell damage. Experiments at earlier time points without cell damage (3h/6h) confirmed that cytokine regulation is independent of mucosal damage.
Monolayer experiments are cheaper than RHE and allow quantitative evaluation of adhesion and invasion. We showed that LGG protects against C. albicans also in monolayer experiments with increased protection after 12h pre-incubation with LGG before addition ofCandida albicans and confirmed that LGG reduces the mucosal immune response in this model. We were also able to show that the presence of LGG strongly reduced the adhesion and invasion ability of C. albicans after 3h. Direct visualisation shows that hyphae of C. albicans are shorter in the presence of LGG and the probiotic reduces hyphal growth of C. albicans by about 40% (figure 3).
Figure 3: LGG Influences Hyphal Growth

Similar protective effects, but immune modulation of different degrees, can be seen with other probiotic Lactobacilli e.g. L. casei shirota (LCS) but this strain does not influence adhesion of C. albicans.
SUMMARY
Using RHE and monolayer cell culture techniques we have shown that L. rhamnosus GG
• Protects mucosa from C. albicans damage in both preventive and therapeutic approaches
• Modulates the immune response
• Induces immune conditioning of mucosal surfaces
Furthermore LGG reduces the adhesion and invasion capacity of C. albicans and inhibits the linear growth of C. albicans hyphae. The ability of such probiotic bacteria to inhibit the growth of pathogens such as C. albicans and to modulate human immune responses to these organisms, offer the promise that these bacteria could provide new options in antifungal therapy.
References
- Schaller M, Zakikhany K, Naglik JR, Weindl G, Hube B. Models of oral and vaginal candidiasis based on in vitro reconstituted human epithelia. Nat Protoc. 2006;1(6):2767-73.
- Mailänder-Sánchez D, Wagener J, Schaller M. Potential role of probiotic bacteria in the treatment and prevention of localised candidosis. Mycoses. 2012 Jan;55(1):17-26.
- Atassi F, Brassart D, Grob P, Graf F, Servin AL. Lactobacillus strains isolated from the vaginal microbiota of healthy women inhibit Prevotella bivia and Gardnerella vaginalis in coculture and cell culture. FEMS Immunol Med Microbiol. 2006 Dec;48(3):424-32
- Manzoni P, Mostert M, Leonessa ML, Priolo C, Farina D, Monetti C, Latino MA, Gomirato G. Oral supplementation with Lactobacillus casei subspecies rhamnosus prevents enteric colonization by Candida species in preterm neonates: a randomized study. Clin Infect Dis. 2006 Jun 15;42(12):1735-42.
- Elahi S, Pang G, Ashman R, Clancy R. Enhanced clearance of Candida albicans from the oral cavities of mice following oral administration of Lactobacillus acidophilus. Clin Exp Immunol. 2005 Jul;141(1):29-36.
PROBIOTICS AND IBS: RATIONALE, PUTATIVE MECHANISM, AND EVIDENCE OF CLINICAL EFFICACY
Professor Robin Spiller, NIHR Nottingham Digestive Diseases Biomedical Research Centre, Nottingham Digestive Diseases Centre, UK.
WHAT IS IRRITABLE BOWEL SYNDROME (IBS)?
Irritable Bowel Syndrome is characterised by recurrent abdominal pain or discomfort associated with disturbed bowel habit. The Rome criteria I, II & III (1992 , 1999 & 2006) were developed to allow uniform entry criteria for randomised clinical trials (RCTs) and to facilitate comparison between studies. The Rome III criteria are summarised in (table 1)1.
Table 1: Rome III criteria for diagnosis of IBS
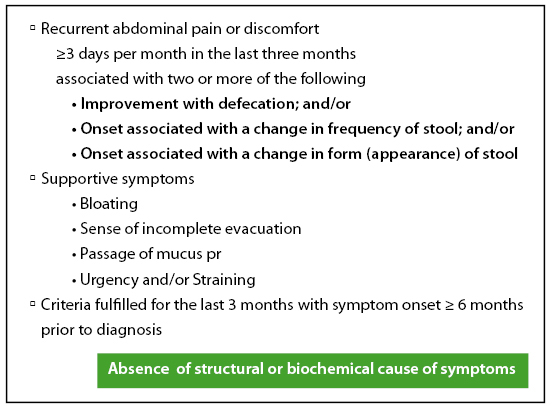
Longstreth et al Gastroenterology 2006;130:1480-91
The time limit for onset of symptoms before diagnosis was designed to avoid giving a chronic disease label to transient symptoms due to e.g. infection. This has been set to 6 months since it is unlikely that a new diagnosis will emerge or that symptoms will disappear if they have already lasted 6 months. Better and faster investigations mean that other diagnoses are now more rapidly eliminated. The threshold of >3 days per month of symptoms is empirical but designed to exclude trivial complaints that might occur at lower frequencies.
IBS is common throughout the world, affecting 10-15% of the adult population (Figure 1).
Figure 1: World wide incidence of IBS
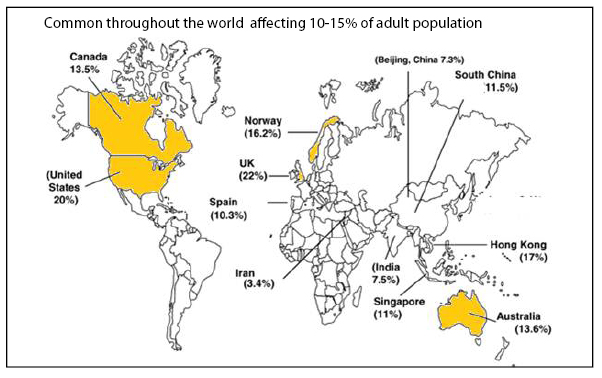
Predictors for the development of IBS include age, gender, infection and stress. Figure 22shows the effect of age and gender on the onset of IBS, and incidentally shows the well known preponderance in females. In both sexes the peak time of onset in the early 20’s corresponds to a common time of stress in people’s lives e.g. associated with leaving home for the first time or starting a first job. This pattern of onset is almost an inverse of the ‘U shaped curve of happiness’ derived from studies of reported wellbeing at different ages3.
Figure 2: Effect of Age and Gender on onset of IBS

Adapted from Jones et al. Aliment Pharmacol Ther 2006; 24(5):879-886
A prospective study in General Practice of 5,250 people free from IBS found 86 new cases in the 2,456 followed up at 15 months4. Gender was the most prominent risk factor with females outnumbering males (4.6% versus 2.1 % respectively OR=2.2). Adjusted for age and sex, other predictors for development of IBS were:
High levels of illness behaviour Odds Ratio (OR) = 5.2 (95% CI 2.5-11.0)
Health anxiety OR = 2.0 (0.98-4.1)
Sleep problems OR = 1.6 (0.8-3.2)
Somatic symptoms OR = 1.6 (0.8-2.9)
Infectious gastroenteritis (IGE) may also be an important predictor of IBS. A meta-analysis5of studies found a sevenfold increase (pooled risk estímate) in the odds of developing IBS following IGE. This post-infectious IBS (PI-IBS) may be susceptible to primary prevention in treating the intial episode of IGE.
Genetic factors are also important. The Virginia twin study6 of 6020 twins found approximately twice the concordance for IBS in monozygotic compared to dizygotic twins (17% v 8.4%). There is some evidence suggesting that Crohn’s associated TNFSF15 polymorphisms are increased in IBS7. This genetic effect should be more obvious in gene/environment interaction and therefore should be most obvious in PI-IBS.
In May 2000, over 2300 residents of Walkerton, Ontario, developed gastroenteritis from microbial contamination of the municipal water supply. A longitudinal study found that 36.2% of these developed IBS8. This allowed researchers to compare genetic data from residents who developed gastroenteritis and reported PI-IBS 2 to 3 years after the outbreak (n = 228, cases), with data from residents who developed gastroenteritis but did not develop PI-IBS (n = 581, controls).
Multivariate analysis after fine mapping of 4 gene areas found significant differences in genes that encode proteins involved in epithelial cell barrier function and the innate immune response to enteric bacteria. These results illustrate the probable importance of host response to bacteria, cytokine response and permeability in PI-IBS.
Figure 3: Risk factors for developing Post Infective IBS after C jejuni enteritis
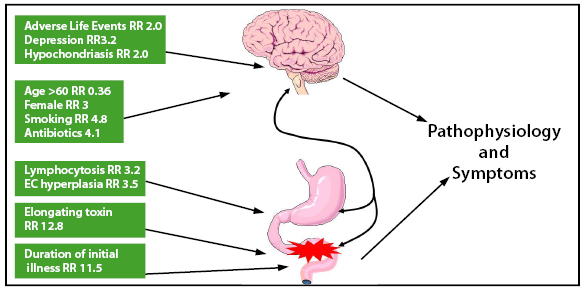
Figure 3 illustrates that most of the known risk factors for PI-IBS reflect local gut damage, but that predisposing psychological factors are also important.
An important role for serotonin or 5-hydroxytryptamine (5HT) in PI-IBS is suggested by evidence that 5-HT containing enterochromaffin (EC) cells increased in PI-IBS but not in those who were infected but recovered fully9, and by data showing that there is increased postprandial plasma 5-HT in PI-IBS10.
Serotonin transporter (SERT) which regulates levels of 5HT may be important in the gut and SERT mRNA appears to be reduced in duodenal biopsies from diarrhoea – predominant IBS (IBS-D)11. There is evidence of low grade ‘immune activation’ in the mucosa of patients with IBS and this may reduce levels of SERT10.
Studies show increased gut permeability in PI-IBS & other IBS subtypes (e.g. IBS-D). Small bowel permeability increased 3 months following Campylobacter enteritis and for up to 4 years in PI-IBS12. This was also seen in PI-IBS 2 years after the Walkerton outbreak13. There is evidence that small bowel permeability is increased in both PI-IBS and IBS-D14.
Increased colonic paracellular permeability has also been seen in IBS patients. Increased permeability resulting from application of the supernatant from IBS colonic biopsies onto cultured cell layers appears to correlate with the severity of abdominal pain15.
Mast cells may provide a link between this increased permeability and pain. Mast cells are known to be potential mediators of the response to stress. Animal studies show psychological stress increases mast cell numbers, and the number of mast cells is increased in IBS16. Moreover IBS colonic biopsies release increased amounts of mast cell products which activate human enteric nerves (histamine, 5-HT & tryptase)17.
ROLE OF ALTERED MICROBIOTA IN DEVELOPING PI-IBS
Acute gastroenteritis leads to depletion of anaerobes, reduction in short chain fatty acids and an increase in pH18. There appears to be an increased risk of developing PI-IBS in Traveller’s Diarrhoea in those treated with antibiotics RR= 4.1(1.1-15.3)19.
SUMMARY
Irritable bowel syndrome is common and ubiquitous.
Risk factors for developing IBS include: female gender; age 20-40 years; infection; and psychological stress possibly acting via mast cells. Abnormal colonic and small bowel permeability is common in IBS. There is evidence of altered microbiota but it is unclear whether this is cause or effect.
RATIONALE FOR USING PROBIOTICS IN IBS
A rationale for the use of probiotics in IBS is given in table 2.
Table 2: Rationale for using probiotics in IBS

ABNORMALITIES OF MICROBIOTA IN IBS
Table 3 shows variable differences in IBS microbiota versus controls from the many studies that have been conducted. There is evidence of increased instability of the microbiota in IBS20, and reduced diversity in IBS-D with reduced Firmicutes/ Bacteroides21.
Table 3: Variable differences in IBS microbiota versus controls

Modified from Simren M, Barbara G, Flint HJ et al. Intestinal microbiota in functional bowel disorders: a Rome foundation report. Gut 2012. Online
CAN WE DRAW ANY CONCLUSIONS?
There are limitations with the available studies including:
• Small numbers of subjects
• With 500 assays we should expect 25 to be different by chance
• Symptoms in IBS are erratic hence analysis at one time point may not be representative – there is a need to assess patients when they are symptomatic.
• Varying methods make comparisons difficult
• Selective cultivation
• Specific targeted qPCR and FISH
• DGGE
• G+C – and microarray profiling
• Patients not well characterized
• Diet, bowel pattern
• >50% of studies come from Finland, and it is uncertain how well these results will generalize to other countries
• Variability a common finding
• Decreased diversity in several diseases with fast transit
• May be more important to link enterotypes or functional characteristics (metabolome) to symptoms rather than individual species
There are clearly many factors that influence the microbiota in IBS (figure 4).
Figure 4: Why might flora be abnormal/unstable in IBS?
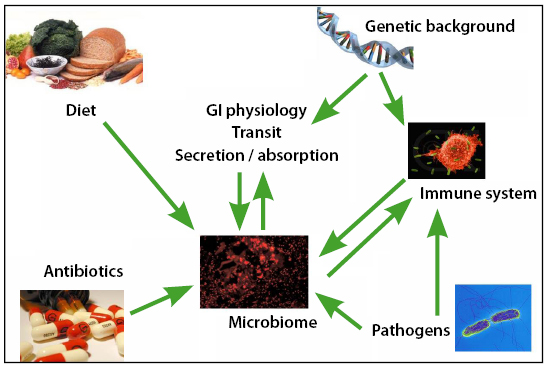
Studies examining whether the gut microbiota are disturbed in IBS have shown a variety of changes which are difficult to interpret. However reasons why microbiota may be disturbed in IBS include:
• Iatrogenic achlorhydria during treatment with proton pump inhibitors (PPIs). Some 44% of IBS sufferers in US tertiary care receive PPIs22.
• Disturbed motility (fast / slow transit). Approximately 1/3 of IBS sufferers have abnormally fast or slow gut transit23 .
• Mucosal immune activation in IBS-D24.
• Depleted diet in attempt to reduce symptoms
• Infectious gastroenteritis leading to post-infectious IBS
• Somatisation leading to frequent doctor visits and increased antibiotic use.
WHICH PROBIOTICS WORK IN IBS?
Tables 4 & 5, and figure 5 summarise the many studies of probiotics in IBS. Those with the most consistent data are circled in red.
Table 4: studies of probiotics in IBS

Table 5: studies of probiotics in IBS
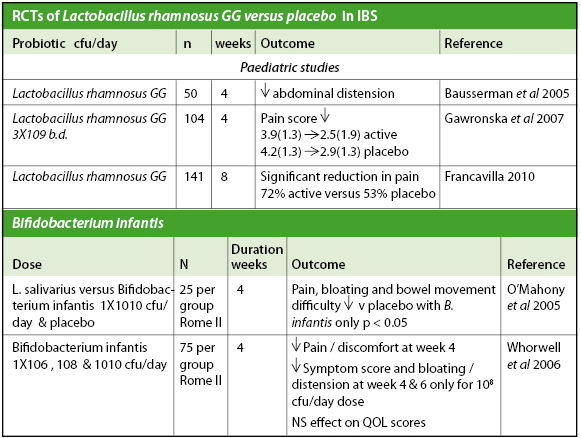
Figure 5: Lactobacillus GG normalises gut permeability in children with IBS & FAP

Reproduced with permission from Pediatrics, Vol. 126, pages e1445-52, Copyright 2010 by the AAP
However there are problems in comparing probiotic studies, including:
1. Variable quality of reporting
2. Variable unvalidated endpoints
3. Use of arbitrary scores rather than numbers of responders
4. Wide range of different species used both single and multiple
5. Meta-analyses abound but are probably invalid except for studies involving the same species.
SUMMARY
IBS patients show evidence of immune activation, impaired gut barrier function and abnormal gut microbiota.
Probiotics can exert anti-inflammatory effects in inflammatory bowel disease and may strengthen the gut barrier in IBS-D25. Randomised Controlled Trials show limited benefit for some, but not all probiotics, with Numbers Needed to Treat varying from 4.8 – infinity, compared to results from conventional pharmacologic agents in IBS of 8 -14. The best evidence of efficacy seems to be for improving pain in children. Defining who will respond remains a challenge.
References
- Longstreth GF, Thompson WG, Chey WD, Houghton LA, Mearin F, Spiller RC. Functional bowel disorders. Gastroenterology 2006 Apr;130(5):1480-91. Review.Erratum in: Gastroenterology. 2006 Aug;131(2):688.
- Jones R, Latinovic R, Charlton J, Gulliford M. Physical and psychological co-morbidity in irritable bowel syndrome: a matched cohort study using the General Practice Research Database. Aliment Pharmacol Ther. 2006 Sep 1;24(5):879-86.
- Stone AA, Schwartz JE, Broderick JE, Deaton A. A snapshot of the age distribution of psychological well-being in the United States. Proc Natl Acad Sci U S A. 2010 Jun 1;107(22):9985-90. Epub 2010 May 17.
- Nicholl BI, Halder SL, Macfarlane GJ, Thompson DG, O’Brien S, Musleh M, McBeth J. Psychosocial risk markers for new onset irritable bowel syndrome--results of a large prospective population-based study. Pain. 2008 Jul;137(1):147-55. Epub 2007 Oct 10.
- Halvorson HA, Schlett CD, Riddle MS. Postinfectious irritable bowel syndrome - a meta-analysis. Am J Gastroenterol. 2006 Aug;101(8):1894-9; quiz 1942.
- Levy RL, Jones KR, Whitehead WE, Feld SI, Talley NJ, Corey LA. Irritable bowel syndrome in twins: heredity and social learning both contribute to etiology. Gastroenterology. 2001 Oct;121(4):799-804.
- Zucchelli M, Camilleri M, Andreasson AN, Bresso F, Dlugosz A, Halfvarson J, Törkvist L, Schmidt PT, Karling P, Ohlsson B, Duerr RH, Simren M, Lindberg G, Agreus L, Carlson P, Zinsmeister AR, D’Amato M. Association of TNFSF15 polymorphism with irritable bowel syndrome. Gut. 2011 Dec;60(12):1671-7.
- Villani AC, Lemire M, Thabane M, Belisle A, Geneau G, Garg AX, Clark WF, Moayyedi P, Collins SM, Franchimont D, Marshall JK. Genetic risk factors for post-infectious irritable bowel syndrome following a waterborne outbreak of gastroenteritis. Gastroenterology. 2010 Apr;138(4):1502-13.
- Dunlop SP, Jenkins D, Neal KR, Spiller RC. Relative importance of enterochromaffin cell hyperplasia, anxiety, and depression in postinfectious IBS. Gastroenterology. 2003 Dec;125 (6):1651-9.
- Foley S, Garsed K, Singh G, Duroudier NP, Swan C, Hall IP, Zaitoun A, Bennett A, Marsden C, Holmes G, Walls A, Spiller RC. Impaired uptake of serotonin by platelets from patients with irritable bowel syndrome correlates with duodenal immune activation. Gastroenterology. 2011 May;140(5):1434-43.
- Dunlop SP, Coleman NS, Blackshaw E, Perkins AC, Singh G, Marsden CA, Spiller RC. Abnormalities of 5-hydroxytryptamine metabolism in irritable bowel syndrome. Clin Gastroenterol Hepatol. 2005 Apr;3(4):349-57.
- Spiller RC, Jenkins D, Thornley JP, Hebden JM, Wright T, Skinner M, Neal KR. Increased rectal mucosal enteroendocrine cells, T lymphocytes, and increased gut permeability following acute Campylobacter enteritis and in post-dysenteric irritable bowel syndrome. Gut. 2000 Dec;47(6):804-11.
- Marshall JK, Thabane M, Garg AX, Clark W, Meddings J, Collins SM; WEL Investigators. Intestinal permeability in patients with irritable bowel syndrome after a waterborne outbreak of acute gastroenteritis in Walkerton, Ontario.Aliment Pharmacol Ther. 2004 Dec;20(11-12):1317-22.
- Dunlop SP, Hebden J, Campbell E, Naesdal J, Olbe L, Perkins AC, Spiller RC. Abnormal intestinal permeability in subgroups of diarrhea-predominant irritable bowel syndromes. Am J Gastroenterol. 2006 Jun;101(6):1288-94. Erratum in: Am J Gastroenterol. 2006 Aug;101(8):1944.
- Piche T, Barbara G, Aubert P, Bruley des Varannes S, Dainese R, Nano JL, Cremon C, Stanghellini V, De Giorgio R, Galmiche JP, Neunlist M. Impaired ntestinal barrier integrity in the colon of patients with irritable bowel syndrome: involvement of soluble mediators. Gut. 2009 Feb;58(2):196-201.
- Barbara G, Stanghellini V, De Giorgio R, Cremon C, Cottrell GS, Santini D,Pasquinelli G, Morselli-Labate AM, Grady EF, Bunnett NW, Collins SM, Corinaldesi R. Activated mast cells in proximity to colonic nerves correlate with abdominal pain in irritable bowel syndrome. Gastroenterology. 2004 Mar;126(3):693-702.
- Buhner S, Li Q, Vignali S, Barbara G, De Giorgio R, Stanghellini V, Cremon C, Zeller F, Langer R, Daniel H, Michel K, Schemann M. Activation of human enteric neurons by supernatants of colonic biopsy specimens from patients with irritable bowel syndrome. Gastroenterology. 2009 Oct;137(4):1425-34. Epub 2009 Jul 28.
- Tazume S, Takeshi K, Saidi SM, Ichoroh CG, Mutua WR, Waiyaki PG, Ozawa A.Ecological studies on intestinal microbial flora of Kenyan children with diarrhoea. J Trop Med Hyg. 1990 Jun;93(3):215-21.
- Stermer E, Lubezky A, Potasman I, Paster E, Lavy A. Is traveler’s diarrhea a significant risk factor for the development of irritable bowel syndrome? A prospective study. Clin Infect Dis. 2006 Oct 1;43(7):898-901. Epub 2006 Aug 25.
- Mättö J, Maunuksela L, Kajander K, Palva A, Korpela R, Kassinen A, Saarela M. Composition and temporal stability of gastrointestinal microbiota in irritable bowel syndrome--a longitudinal study in IBS and control subjects. FEMS Immunol Med Microbiol. 2005 Feb 1;43(2):213-22.
- Rajilic-Stojanovic M, Biagi E, Heilig HG, Kajander K, Kekkonen RA, Tims S, de Vos WM. Global and deep molecular analysis of microbiota signatures in fecal samples from patients with irritable bowel syndrome. Gastroenterology. 2011 Nov;141(5):1792-801. Epub 2011 Aug 5.
- Majewski M, McCallum RW. Results of small intestinal bacterial overgrowth testing in irritable bowel syndrome patients: clinical profiles and effects of antibiotic trial. Adv Med Sci. 2007;52:139-42.
- Camilleri M, McKinzie S, Busciglio I, Low PA, Sweetser S, Burton D, Baxter K, Ryks M, Zinsmeister AR. Prospective study of motor, sensory, psychologic, and autonomic functions in patients with irritable bowel syndrome. Clin Gastroenterol Hepatol. 2008 Jul;6(7):772-81. Epub 2008 May 5.
- Chadwick VS, Chen W, Shu D, Paulus B, Bethwaite P, Tie A, Wilson I. Activation of the mucosal immune system in irritable bowel syndrome. Gastroenterology. 2002 Jun;122(7):1778-83.
- Zeng J, Li YQ, Zuo XL, Zhen YB, Yang J, Liu CH. Clinical trial: effect of active lactic acid bacteria on mucosal barrier function in patients with diarrhoea-predominant irritable bowel syndrome. Aliment Pharmacol Ther. 2008 Oct 15;28(8):994-1002.
TRENDS IN PROBIOTICS: TARGETS AND MECHANISMS OF ACTION
Dr Cormac G. Gahan, Alimentary Pharmabiotic Centre & Department of Microbiology, University College Cork, Ireland.
Intestinal bacteria can be thought of as a hidden organ. The intestines contain more bacteria (representing approximately 2000 phylotypes) than cells in the human body, with total metabolic activity rivalling that of the liver. It is clear that the gut microbiota can be both an asset and a liability:
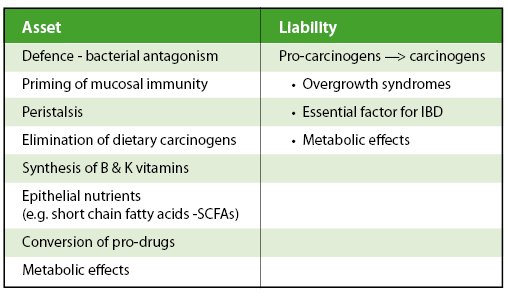
The Alimentary Pharmabiotic Centre in Cork has a multidisciplinary team working in state of the art facilities to ‘mine microbes for mankind’. Figure 1 outlines the research approach.
Figure 1: APC research outline

Recent Highlights of APC Research include:
MAPPING THE GUT MICROBIOTA IN THE ELDERLY1
Alterations in the human intestinal microbiota are linked to conditions including inflammatory bowel disease, irritable bowel syndrome, and obesity. The microbiota also undergoes substantial changes at the extremes of life, in infants and older people, the ramifications of which are still being explored.
Claesson and co-workers applied pyrosequencing to characterize the faecal microbiota in 161 subjects aged 65y and older and 9 younger control subjects. The core microbiota of elderly subjects was distinct from that previously established for younger adults, with a greater proportion of Bacteroides spp. and distinct abundance patterns of Clostridium groups. Analyses of 26 faecal microbiota datasets from 3-month follow-up samples indicated that in 85% of the subjects, the microbiota composition was more like the corresponding time-0 sample than any other dataset. The group concluded that the faecal microbiota of the elderly shows temporal stability over limited time in the majority of subjects but is characterized by unusual phylum proportions and extreme variability (Figure 2).
Figure 2:
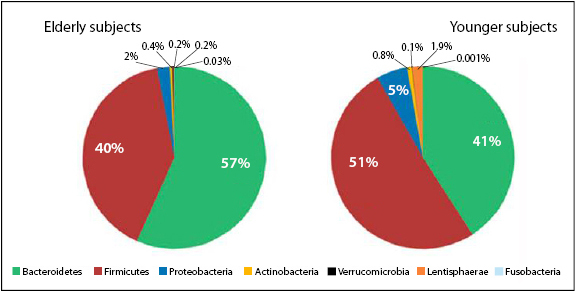
Claesson M J et al. PNAS 2011;108:4586-4591
STUDIES ON MECHANISMS OF COLONISATION BY BIFIDOBACTERIUM BREVE2
Development of the human gut microbiota commences at birth, with bifidobacteria being among the first colonizers of the sterile newborn gastrointestinal tract. To date, the genetic basis of Bifidobacterium colonization and persistence remains poorly understood.
Mutational analysis demonstrated that the tad(2003) gene cluster is essential for efficient in vivo murine gut colonization, and immunogold transmission electron microscopy confirmed the presence of Tad pili at the poles of B. breve UCC2003 cells (Figure 3).
Figure 3: Identification of pili in B. breve UCC2003 by immunogold electron microscopy

O’Connell Motherway M et al. PNAS 2011;108:11217-11222
Conservation of the Tad pilus-encoding locus among other B. breve strains and among sequenced Bifidobacterium genomes supports the notion of a ubiquitous pili-mediated host colonization and persistence mechanism for bifidobacteria.
WORK SHOWING THAT L. RHAMNOSUS (JB-1) REDUCES STRESS-INDUCED ANXIETY- AND DEPRESSION-RELATED BEHAVIOR IN MICE3
There is increasing, but largely indirect, evidence pointing to an effect of commensal gut microbiota on the central nervous system (CNS). However, it is unknown whether lactic acid bacteria such as Lactobacillus rhamnosus could have a direct effect on neurotransmitter receptors in the CNS in normal, healthy animals.
In this work, Bravo et al. showed that chronic treatment with L. rhamnosus (JB-1) induced region-dependent alterations in GABA mRNA expression in the brain, in comparison with control-fed mice. Importantly, L. rhamnosus (JB-1) reduced stress-induced corticosterone and anxiety- and depression-related behavior. Moreover, the neurochemical and behavioral effects were not found in vagotomized mice, identifying the vagus as a major modulatory constitutive communication pathway between the bacteria exposed to the gut and the brain.
PROBIOTICS AS ANTI-INFECTIVES – MECHANISMS4
In recent years, there has been a growing interest in the use of probiotic bacteria for the maintenance of general gastrointestinal health and the prevention or treatment of intestinal infections. Whilst probiotics are documented to reduce or prevent specific infectious diseases of the GI tract, the mechanistic basis of this effect remains unclear.
It is likely that diverse modes-of-action contribute to inhibition of pathogens in the gut environment and proposed mechanisms include (i) direct antimicrobial activity through production of bacteriocins or inhibitors of virulence gene expression; (ii) competitive exclusion by competition for binding sites or stimulation of epithelial barrier function; (iii) stimulation of immune responses via increases of sIgA and anti-inflammatory cytokines and regulation of proinflammatory cytokines; and (iv) inhibition of virulence gene or protein expression in gastrointestinal pathogens.
There is evidence for a role of probiotics both in prevention of infection (e.g. diarrhoea in children5) and in prevention of antibiotic associated diarrhoea6 (Figure 4).
Figure 4: Proposed Mechanisms

In vitro studies, support the hypothesis that the ability of probiotic agents to inhibit adherence of organisms to intestinal epithelial cells is mediated through their ability to increase expression of MUC2 and MUC3 intestinal mucins7.
The microbiota fosters development, aids digestion and protects host cells from pathogens – a function referred to as colonization resistance. Gut inflammation changes microbiota composition, disrupts colonization resistance and enhances pathogen growth. Thus, some pathogens can benefit from inflammatory defenses8.
FOCUS UPON BACTERIOCIN PRODUCTION
Workers from the APC have showed that Lactobacillus salivarius UCC118, a recently sequenced and genetically tractable probiotic strain of human origin, produces a bacteriocinin vivo that can significantly protect mice against infection with the invasive foodborne pathogen Listeria monocytogenes9. This raises the intriguing possibility of probiotics specifically designed to protect against infection in particular circumstances such as pregnancy.
CHARACTERISATION OF THURICIN
Another bacteriocin of interest in the APC is Thuricin CD a bacteriocin produced by Bacillus thuringiensis. Thuricin CD is a two component bacteriocin that is in the drug discovery pipeline as a candidate against the target pathogen C. difficile10 (Figure 5).
Figure 5: Colony of B. thuringiensis DPC 6431 on the initial isolation plate showing inhibition in seeded overlay of C. difficile ATCC 43593

Reproduced with permission from PNAS. Rea M C et al. PNAS 2010;107:9352-9357
This bacteriocin offers the promise of a narrow-spectrum antimicrobial with specific anti-C. difficile activity. Thuricin CD was effective at killing C. difficile in the distal colon model but had no significant impact on the composition of the microbiota. This offers the possibility of developing a targeted approach to eliminating C. difficile in the colon, without collateral damage11.
Figure 6: Minimal alterations to background microbiota following Thuricin treatment
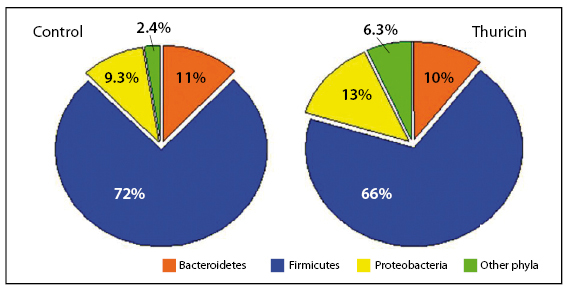
Reproduced with permission from PNAS. Rea M C et al. PNAS 2011;108:4639-4644
STUDIES OF BILE SALT HYDOLASE (BSH) AS A COMMON GUT-SPECIFIC BACTERIAL FUNCTION
BSH activity is a conserved microbial adaptation to the human gut environment with a high level of redundancy in this ecosystem. Through metagenomic analyses we have identified functional BSH in all major bacterial divisions and archaeal species in the gut and demonstrated that BSH is enriched in the human gut microbiome.
Phylogenetic analysis illustrates that selective pressure in the form of conjugated bile acid has driven the evolution of members of the Ntn_CGH-like family of proteins toward BSH activity in gut-associated species. Furthermore, we have demonstrated that BSH mediates bile tolerance in vitro and enhances survival in the murine gut in vivo12.
Reabsorbed bile salts act as signalling molecules regulating systemic endocrine functions13and bile acids act as local signalling molecules regulating innate immunity14. Regulatory networks recognising bile acids are affected by the microbiota15.
We can conclude that functional metagenomic analysis has helped to define the Bile Salt Hydrolase complement in humans. Functional BSH is distributed across gut Bacteria andArchaea. Bile Salt Hydrolase is a widespread gut specific trait and may have implications for overall health in the host. Overall the microbiota alters bile acid recirculation and regulatory networks recognising bile acids are therefore affected by the microbiota. Since bile acids have important roles as signalling molecules and in regulating inate immunity these findings have considerable potential for eventual therapeutic applications.
SUMMARY
I have given a snapshot of some of the important work going on at the APC. The creation of a state of the art facility, and a multidisciplinary team with extensive international collaborations and the exciting work ongoing, sends a clear message that the alimentary microbiota is a source of great potential as we ‘mine microbes for mankind’.
References
- Claesson MJ, Cusack S, O’Sullivan O, Greene-Diniz R, de Weerd H, Flannery E,Marchesi JR, Falush D, Dinan T, Fitzgerald G, Stanton C, van Sinderen D, O’Connor M, Harnedy N, O’Connor K, Henry C, O’Mahony D, Fitzgerald AP, Shanahan F, Twomey C, Hill C, Ross RP, O’Toole PW. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc Natl Acad Sci U S A. 2011 Mar 15;108 Suppl 1:4586-91. Epub 2010 Jun 22.
- O’Connell Motherway M, Zomer A, Leahy SC, Reunanen J, Bottacini F, Claesson MJ, O’Brien F, Flynn K, Casey PG, Munoz JA, Kearney B, Houston AM, O’Mahony C, Higgins DG, Shanahan F, Palva A, de Vos WM, Fitzgerald GF, Ventura M, O’Toole PW, van Sinderen D. Functional genome analysis of Bifidobacterium breve UCC2003 reveals type IVb tight adherence (Tad) pili as an essential and conserved host-colonization factor. Proc Natl Acad Sci U S A. 2011 Jul 5;108(27):11217-22. Epub 2011 Jun 20.
- Bravo JA, Forsythe P, Chew MV, Escaravage E, Savignac HM, Dinan TG, Bienenstock J, Cryan JF. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc Natl Acad Sci U S A. 2011 Sep 20;108(38):16050-5. Epub 2011 Aug 29.
- Corr SC, Hill C, Gahan CG. Understanding the mechanisms by which probiotics inhibit gastrointestinal pathogens. Adv Food Nutr Res. 2009;56:1-15. Review.
- Sur D, Manna B, Niyogi SK, Ramamurthy T, Palit A, Nomoto K, Takahashi T, Shima T, Tsuji H, Kurakawa T, Takeda Y, Nair GB, Bhattacharya SK. Role of probiotic in preventing acute diarrhoea in children: a community-based, randomized, double-blind placebo-controlled field trial in an urban slum. Epidemiol Infect. 2011 Jun;139(6):919-26. Epub 2010 Jul 30.
- Gao XW, Mubasher M, Fang CY, Reifer C, Miller LE. Dose-response efficacy of a proprietary probiotic formula of Lactobacillus acidophilus CL1285 and Lactobacillus casei LBC80R for antibiotic-associated diarrhea and Clostridium difficile-associated diarrhea prophylaxis in adult patients. Am J Gastroenterol. 2010 Jul;105(7):1636-41. Epub 2010 Feb 9.
- Mack DR, Michail S, Wei S, McDougall L, Hollingsworth MA. Probiotics inhibit enteropathogenic E. coli adherence in vitro by inducing intestinal mucin gene expression. Am J Physiol. 1999 Apr;276(4 Pt 1):G941-50.
- Stecher B, Hardt WD. The role of microbiota in infectious disease. Trends Microbiol. 2008 Mar;16(3):107-14. Epub 2008 Feb 14. Review.
- Corr SC, Li Y, Riedel CU, O’Toole PW, Hill C, Gahan CG. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc Natl Acad Sci U S A. 2007 May 1;104(18):7617-21. Epub 2007 Apr 24.
- Rea MC, Sit CS, Clayton E, O’Connor PM, Whittal RM, Zheng J, Vederas JC, Ross RP, Hill C. Thuricin CD, a posttranslationally modified bacteriocin with a narrow spectrum of activity against Clostridium difficile. Proc Natl Acad Sci U S A. 2010 May 18;107(20):9352-7. Epub 2010 Apr 30.
- Rea MC, Dobson A, O’Sullivan O, Crispie F, Fouhy F, Cotter PD, Shanahan F, Kiely B, Hill C, Ross RP. Effect of broad- and narrow-spectrum antimicrobials on Clostridium difficile and microbial diversity in a model of the distal colon. Proc Natl Acad Sci U S A. 2011 Mar 15;108 Suppl 1:4639-44. Epub 2010 Jun 29.
- Jones BV, Begley M, Hill C, Gahan CG, Marchesi JR. Functional and comparative metagenomic analysis of bile salt hydrolase activity in the human gut microbiome. Proc Natl Acad Sci U S A. 2008 Sep 9;105(36):13580-5. Epub 2008 Aug 29.
- Watanabe M, Houten SM, Mataki C, Christoffolete MA, Kim BW, Sato H, Messaddeq N, Harney JW, Ezaki O, Kodama T, Schoonjans K, Bianco AC, Auwerx J. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature. 2006 Jan 26;439(7075):484-9. Epub 2006 Jan 8.
- Inagaki T, Moschetta A, Lee YK, Peng L, Zhao G, Downes M, Yu RT, Shelton JM, Richardson JA, Repa JJ, Mangelsdorf DJ, Kliewer SA. Regulation of antibacterial defense in the small intestine by the nuclear bile acid receptor. Proc Natl Acad Sci U S A. 2006 Mar 7;103(10):3920-5. Epub 2006 Feb 10.
- Swann JR, Want EJ, Geier FM, Spagou K, Wilson ID, Sidaway JE, Nicholson JK, Holmes E. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc Natl Acad Sci U S A. 2011 Mar 15;108 Suppl 1:4523-30. Epub 2010 Sep 13.
PROBIOTICS, PREBIOTICS AND SYNBIOTICS: APPROACHES FOR MODULATING THE MICROBIAL ECOLOGY OF THE GUT
Professor Ian Rowland, Department of Food & Nutritional Sciences University of Reading, UK.
Probiotics, often in foods such as yogurt or yogurt drinks, are now promoted and consumed to promote intestinal health. Important to assessing whether this is an effective strategy, is knowing whether these organisms:
• Survive in the gut
• Stimulate other lactic acid bacteria (LAB)
• Inhibit pathogens
There are some data to support each of these factors influencing the activity of such agents. In most studies probiotic consumption usually results in temporary ‘colonization’ of the gut by the administered strain. A study in 20 healthy volunteers showed that daily consumption of a fermented milk drink (~1011 cfu L. casei per day for 21 days) enabled the probiotic Lact. casei strain to be maintained in the gastrointestinal tract (recovered from faecal samples) at a stable relatively high population level during the probiotic feeding period1.
Studies, including in infants and elderly subjects also confirm stimulation of other LAB species in the gut – consumption of lactobacilli stimulates bifids and vice versa2,3. Some studies have also shown a decrease in potential pathogens4,5,6. One randomised double blind placebo controlled study with L. casei taken during and one week after a course of antibiotics, showed that it provided significant protection against antibiotic associated diarrhea and significantly reduced the presence of C. difficile toxin7.
More complex and sensitive molecular methodologies reveal extensive changes in LAB and also unrelated bacterial phylotypes after probiotic consumption.
PREBIOTIC DEFINITION
The International Scientific Association for Probiotics and Prebiotics8 (2008, London, Ontario) agreed the following definition for a prebiotic:
‘A dietary prebiotic is a selectively fermented ingredient that results in specific changes, in the composition and/or activity of the gastrointestinal microbiota, thus conferring benefit(s) upon host health.’
Dietary components that reach the colon, and are available to influence the microbiota include poorly digestible carbohydrates, such as non-starch polysaccharides, resistant starch, non-digestible oligosaccharides (NDOs) and polyphenols.
Resistant Starch is classified in the following way (with examples of each type):
RS1: Physically inaccessible starch
• partly milled grains and seeds
RS2: Native starch granule
• raw potato and green banana
RS3: Retrograded starch (mainly amylose)
• cooled-cooked potato, bread, corn flakes
RS4: Chemically modified (cross-linking, esterification)
RS1, in the form of whole grain cereal has been shown to have more impact in increasing lactobacilli compared to wheat bran alone9. RS3 has been shown to modify gut microbiota in rats, greatly increasing populations of bifidobacteria and lactobacilli and reducing populations of potentially pathogenic bacterial groups10.
NON-DIGESTIBLE OLIGOSACCHARIDES (NDOS)
Non-digestible oligosaccharides occur in plants (chicory, asparagus, artichoke, onions, garlic, leeks, soy beans) and human breast milk. NDOs have an osidic bond resistant to hydrolysis by intestinal digestive enzymes and this means that they are poorly digested in the upper gastrointestinal tract and reach the colon intact. There they are fermented by colonic microflora to produce short chain fatty acids (SCFA) and gas. NDO substances are widely used in many food and drink products, including yoghurts. Categories of NDO are given in Table 1:
Table 1: Categories of non-digestible oligosaccharides
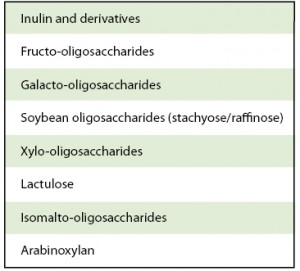
A number of oligosaccharides have been reported to be prebiotic, but the quality of the data is variable. Those with the best evidence include: inulin, fructo-oligosaccharides, galacto-oligosaccharides and lactulose, which have all been shown to have ‘bifidogenic’ effects. The effects of NDO on gut microbiota are summarised in Table 2.
Table 2: effects of NDOs on gut microbiota:
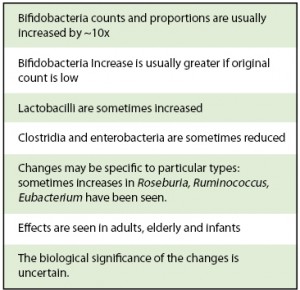

Beyond the usual categories of prebiotic substances a range of other plant derived substances such as certain flavonoids notably catechins have the potential to influence gut microbiota11. There is some evidence that consumption of cocoa flavonoids influences the composition of gut bacteria in vivo12.
SYNBIOTICS
Synbiotics have been defined as:
‘A mixture of probiotics and prebiotics that beneficially affects the host by improving the survival and implantation of live microbial dietary supplements in the gastrointestinal tract, by selectively stimulating the growth and/or activating the metabolism of one or a limited number of health-promoting bacteria, and thus improving welfare.’13
Combining probiotics and prebiotics is potentially a very effective way of influencing gut microbiota and health14 but as yet there are few in vivo studies in humans; and none that compare pro, pre and synbiotics directly.
GUT MICROBIOTA – WHAT SHOULD WE MEASURE?
The majority of studies that have examined the effects of probiotics and prebiotics on the gut microflora have focused on changes in bifidobacteria, but this raises the question: Can increased bifidobacteria alone be considered as a health benefit?
This question gives rise to a number of subsidiary questions including:
• Are bifidogenic probiotics associated with health benefits?
• Are high bifidobacteria associated with positive health?
• Are low faecal bifidobacteria associated with poorer health?
• Do prebiotics always increase bifidobacteria in all subjects?
• Are prebiotic health benefits always associated with increased bifidobacteria?
• How big an increase in bifidobacteria is needed for a health benefit?
• Is bifidogenic effect a biomarker of other changes in the microflora?
Figure 1: Claimed Health Benefits of Prebiotics
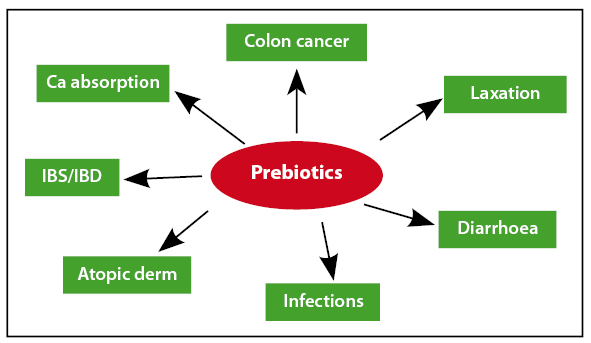
Figure 1 summarises some of the health benefits claimed for prebiotics. An enormous amount of work has been done and continues to be done to link prebiotic health benefits and changes in bifidobacteria.
The current evidence that claimed health benefits for prebiotics are associated with increased bifidobacteria is summarized in Table 3:
Table 3: Status of Evidence for Prebiotic Health Benefits being associated with changes in bifidobacteria numbers

CONCLUSIONS
We can show that probiotics, prebiotics and synbiotics influence gut microbial ecology. Their main effect is to increase bifidobacteria and lactobacilli, but metagenomic approaches may reveal wider effects. Changes in the microbiota are often associated with health or physiological benefits. It is clear that some plant polyphenols may act as prebiotics, providing a partial explanation for some of the health benefits associated with high dietary consumption of vegetables.
Research questions for the future include:
• Should we be searching for probiotics other than bifidobacteria and lactobacilli?
• Should we develop more species-targeted prebiotics/synbiotics?
• Should we be less focused on LAB changes as indicators of prebiotic effects and include other possible indicators e.g. Roseburia, F.prausnitzii, C.butyricum?
• Is it important to consider more holistic changes in microbiota – e.g. changes in diversity?
References
- Tuohy KM, Pinart-Gilberga M, Jones M, Hoyles L, McCartney AL, Gibson GR. Survivability of a probiotic Lactobacillus casei in the gastrointestinal tract of healthy human volunteers and its impact on the faecal microflora. J Appl Microbiol. 2007 Apr;102(4):1026-32.
- Marzotto M, Maffeis C, Paternoster T, Ferrario R, Rizzotti L, Pellegrino M, Dellaglio F, Torriani S. Lactobacillus paracasei A survives gastrointestinal passage and affects the fecal microbiota of healthy infants. Res Microbiol. 2006 Nov;157(9):857-66. Epub 2006 Aug 2.
- Lahtinen SJ, Tammela L, Korpela J, Parhiala R, Ahokoski H, Mykkänen H, Salminen SJ. Probiotics modulate the Bifidobacterium microbiota of elderly nursing home residents. Age (Dordr). 2009 Mar;31(1):59-66. Epub 2008 Nov 28.
- Manzoni P. Use of Lactobacillus casei subspecies Rhamnosus GG and gastrointestinal colonization by Candida species in preterm neonates. J Pediatr Gastroenterol Nutr. 2007 Dec;45 Suppl 3:S190-4. Review. Erratum in: J Pediatr Gastroenterol Nutr. 2009 Jan;48(1):121-2.
- Lahtinen SJ, Forssten S, Aakko J, Granlund L, Rautonen N, Salminen S, Viitanen M, Ouwehand AC. Probiotic cheese containing Lactobacillus rhamnosus HN001 and Lactobacillus acidophilus NCFM® modifies subpopulations of fecal lactobacilli and Clostridium difficile in the elderly. Age (Dordr). 2012 Feb;34(1):133-43. Epub 2011 Jan 25.
- Szachta P, Ignys I, Cichy W. An evaluation of the ability of the probiotic strain Lactobacillus rhamnosus GG to eliminate the gastrointestinal carrier state of vancomycin-resistant enterococci in colonized children. J Clin Gastroenterol. 2011 Nov-Dec;45(10):872-7.
- Hickson M, D’Souza AL, Muthu N, Rogers TR, Want S, Rajkumar C, Bulpitt CJ. Use of probiotic Lactobacillus preparation to prevent diarrhoea associated with antibiotics: randomised double blind placebo controlled trial. BMJ. 2007 Jul 14;335(7610):80. Epub 2007 Jun 29.
- International Scientific Association for Probiotics and Prebiotics, Nov 2008, London, Ontario. . Last accessed June 2012
Reference Link - Carvalho-Wells AL, Helmolz K, Nodet C, Molzer C, Leonard C, McKevith B, Thielecke F, Jackson KG, Tuohy KM. Determination of the in vivo prebiotic potential of a maize-based whole grain breakfast cereal: a human feeding study. Br J Nutr. 2010 Nov;104(9):1353-6.
- Kleessen B, Stoof G, Proll J, Schmiedl D, Noack J, Blaut M. Feeding resistant starch affects fecal and cecal microflora and short-chain fatty acids in rats. J Anim Sci. 1997 Sep;75(9):2453-62.
- Tzounis X, Vulevic J, Kuhnle GG, George T, Leonczak J, Gibson GR, Kwik-Uribe C, Spencer JP. Flavanol monomer-induced changes to the human faecal microflora. Br J Nutr. 2008 Apr;99(4):782-92.
- Tzounis X, Rodriguez-Mateos A, Vulevic J, Gibson GR, Kwik-Uribe C, Spencer JP. Prebiotic evaluation of cocoa-derived flavanols in healthy humans by using a randomized, controlled, double-blind, crossover intervention study. Am J Clin Nutr. 2011 Jan;93(1):62-72.
- Gibson GR, Roberfroid MB. Dietary modulation of the human colonic microbiota: introducing the concept of prebiotics. J Nutr. 1995 Jun;125(6):1401-12. Review.
- Saulnier DM, Gibson GR, Kolida S. In vitro effects of selected synbiotics on the human faecal microbiota composition. FEMS Microbiol Ecol. 2008 Dec;66(3):516-27.
- Moro G, Arslanoglu S, Stahl B, Jelinek J, Wahn U, Boehm G. A mixture of prebiotic oligosaccharides reduces the incidence of atopic dermatitis during the first six months of age. Arch Dis Child. 2006 Oct;91(10):814-9.
- Arslanoglu S, Moro GE, Schmitt J, Tandoi L, Rizzardi S, Boehm G. Early dietary intervention with a mixture of prebiotic oligosaccharides reduces the incidence of allergic manifestations and infections during the first two years of life. J Nutr. 2008 Jun;138(6):1091-5.
- Arslanoglu S, Moro GE, Boehm G. Early supplementation of prebiotic oligosaccharides protects formula-fed infants against infections during the first 6 months of life. J Nutr. 2007 Nov;137(11):2420-4.
- Cummings JH, Christie S, Cole TJ. A study of fructo oligosaccharides in the prevention of travellers’ diarrhoea. Aliment Pharmacol Ther. 2001 Aug;15(8):1139-45.
- Drakoularakou A, Tzortzis G, Rastall RA, Gibson GR. A double-blind, placebo-controlled, randomized human study assessing the capacity of a novel galacto-oligosaccharide mixture in reducing travellers’ diarrhoea. Eur J Clin Nutr. 2010 Feb;64(2):146-52. Epub 2009 Sep 16.
- Lewis S, Burmeister S, Cohen S, Brazier J, Awasthi A. Failure of dietary oligofructose to prevent antibiotic-associated diarrhoea. Aliment Pharmacol Ther. 2005 Feb 15;21(4):469-77
- Lewis S, Burmeister S, Brazier J. Effect of the prebiotic oligofructose on relapse of Clostridium difficile-associated diarrhea: a randomized, controlled study. Clin Gastroenterol Hepatol. 2005 May;3(5):442-8.
- Abrams SA, Griffin IJ, Hawthorne KM, Liang L, Gunn SK, Darlington G, Ellis KJ. A combination of prebiotic short- and long-chain inulin-type fructans enhances calcium absorption and bone mineralization in young adolescents. Am J Clin Nutr. 2005 Aug;82(2):471-6.
- Heilpern D, Szilagyi A. Manipulation of intestinal microbial flora for therapeutic benefit in inflammatory bowel diseases: review of clinical trials of probiotics, pre-biotics and synbiotics. Rev Recent Clin Trials. 2008 Sep;3(3):167-84. Review.
- Silk DB, Davis A, Vulevic J, Tzortzis G, Gibson GR. Clinical trial: the effects of a trans-galactooligosaccharide prebiotic on faecal microbiota and symptoms in irritable bowel syndrome. Aliment Pharmacol Ther. 2009 Mar 1;29(5):508-18. Epub 2008 Dec 2.
